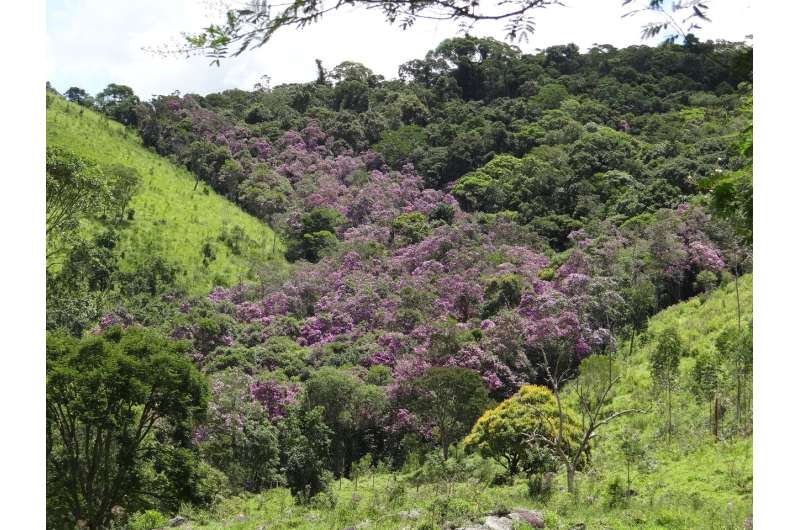
Tibouchina pulchra (glory bush) clearly indicates forest regeneration as almost all the individuals of this species were located inside or adjacent to forest remnants that have recovered since 1962. Credit: Fabien Hubert Wagner
The Atlantic Rainforest has been so savagely clearcut and burned over several centuries that only approximately 12% now remains. Nevertheless, it is still one of the planet's largest repositories of biodiversity, and counter to a process that appeared irreversible, forest cover in the biome has begun to recover in recent decades.
To confirm this trend and understand the dynamics of forest fragment degradation and regeneration, a study conducted at the National Space Research (INPE) in Brazil compared recent high-resolution satellite images with georeferenced aerial photographs taken in 1962 and deployed powerful computational resources to analyze the changes in forest cover using two pioneer tree species as markers: Cecropia hololeuca (silver embaúba) and Tibouchina pulchra (glory bush).
Pioneer species are hardy species that are the first to colonize previously disrupted or damaged ecosystems. An article on the study is published in the journal PLOS ONE.
"Both species are pioneers that grow after degradation. Our study shows that C. hololeuca is a marker of degraded forest and that T. pulchra is an indicator of forest regeneration on former pastureland. By comparing the images, we were able to determine whether the forest fragments investigated evolved from pasture or were already there before 1962," Fabien Hubert Wagner, lead author of the article, told.
Wagner explained that understanding the history of Atlantic Rainforest remnants is important because biodiversity is potentially greatest in older, less degraded forest formations, and this kind of information is crucial for conservation.
The region studied is in the state of São Paulo and lies between São José dos Campos, a large upland city, and Ubatuba on the coast. It includes part of the Serra do Mar State Park. "There are over 2,000 forest fragments in this quadrilateral, which measure approximately 100 km by 60 km," Wagner said. "We computed maps of species dominance for these two tree species and found that at least 4.3% of the region's natural forest cover regenerated after 1962."
Wagner believes these areas regenerated after the passage of Federal Law 5106 in 1966, whereby afforestation or reforestation initiatives were encouraged by means of tax incentives. "Another interesting finding of the study that corroborates this hypothesis is that the eucalyptus groves seen in the region today were planted where there was previously no forest cover," he said. "They're poor in biodiversity but contributed to the expansion of forest cover."
In addition to the excellent database comprising images produced by the WorldView-2 and WorldView-3 satellites, with spatial resolutions of 0.5 m and 0.3 m, respectively, as well as 40 aerial photographs taken in 1962, the study benefited from the use of the powerful artificial intelligence/machine learning tool U-net, a convolutional neural network (CNN).
"It reproduces what the human eye does but on an incomparably larger scale. It can map millions of objects in a single image," Wagner said.
Convolutional neural networks are deep learning algorithms used for image classification and recognition because of their high accuracy.
"We found that T. pulchra clearly indicates forest regeneration, as almost all the individuals of this species were located inside or adjacent to forest remnants that have recovered since 1962. C. hololeuca, however, appeared almost exclusively in older fragments already present in 1962," Wagner said.
The same procedure will be extended to the entire Atlantic Rainforest biome, remnants of which are located on the coast of 17 Brazilian states. "It isn't enough to know there's a forest in a particular area. We need to know its history, whether it's old or recent, in order to conserve it. That's the aim of our research," Wagner said.
More information: Fabien H. Wagner et al, Mapping Atlantic rainforest degradation and regeneration history with indicator species using convolutional network, PLOS ONE (2020). DOI: 10.1371/journal.pone.0229448
Journal information: PLoS ONE
Provided by FAPESP