Distinct processing of lncRNAs contributes to non-conserved functions in stem cells
Long noncoding RNAs (lncRNAs), which are longer than 200 nucleotides in length and lack protein coding potential, are pervasively transcribed in eukaryotic genomes. It is well established that lncRNAs play important roles in gene expression in diverse cellular and biological progress.
Unlike conserved sequences of messenger RNAs (mRNAs), lncRNAs in general lack high constraint of primary sequence and evolve more rapidly than mRNAs. Conservation of lncRNAs, therefore, occurs at different levels from primary sequence, exonic structure, genomic position to mechanism of action. Whether conserved lncRNAs undergo conserved processing, localization and function has remained unexplored.
A research team led by Dr. CHEN Lingling from Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences, discovered that distinct processing of lncRNA orthologs resulted in different subcellular localization in human and murine embryonic stem cells (ESCs). This subsequently leads to their functional divergence in the context of pluripotency regulation across species.
These findings, published in Cell, suggested that conserved lncRNAs may achieve functional evolution through non-conserved RNA processing and localization.
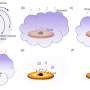
Through the examination of cytoplasmic and nuclear fractionated RNAs of human and murine ESCs, the researchers revealed different subcellular localization patterns of sequence-conserved and positionally conserved lncRNAs in these different cells, and a significantly higher fraction of lncRNAs is more frequently spliced and exported to the cytoplasm in human ESCs than in murine ESCs. This turns out to be important for human ESC pluripotency.
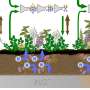
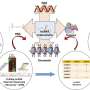
FOXD3 antisense transcript 1(FAST) is a positionally conserved lncRNA but it not conserved in its processing and localization. In human ESCs, the cytoplasm-localized hFAST binds to the WD40 repeat domain of the E3 ubiquitin ligase β-TrCP to prevent β-catenin from degradation, leading to activated WNT signaling required for human ESC pluripotency. Whereas mFast is nuclear retained in murine ESCs and is not essential for self-renewal.
The researchers further uncovered a key processing factor Peptidylprolyl Isomerase E (PPIE) that is responsible for such distinct processing of lncRNAs across species. PPIE is highly expressed in murine ESCs and suppresses processing and nuclear export of many lncRNAs, including mFast. In contrast, the low expression of PPIE in human and monkey ESCs enables hFAST to be efficiently spliced and exported to the cytoplasm, where hFAST acts to promote stem cell self-renewal.
Collectively, this study highlights a mechanism of rapid lncRNA evolution and that the relatively high evolutionary plasticity of lncRNAs can support species-specific gene expression programs.
More information: Chun-Jie Guo et al. Distinct Processing of lncRNAs Contributes to Non-conserved Functions in Stem Cells, Cell (2020). DOI: 10.1016/j.cell.2020.03.006
Journal information: Cell
Provided by Chinese Academy of Sciences