Physically producing computer-generated artificial genomes to understand DNA
The molecular blueprint of life is stored in DNA within the genome. The digital revolution in biology, driven by DNA sequencing, enables scientists to read the genomes of the many microbes and multicellular organisms that populate our world. Today, DNA sequences of over 200,000 microbial genomes are deposited in digital genome databases and have exponentially increased the understanding of how DNA programs living systems. Using this incredible treasure trove of molecular building blocks, bioengineers learn to sequence and synthesize long DNA molecules and to breed useful microbes with the help of computers.
In his research, Beat Christen, professor of experimental systems biology, and researchers in the Christen Lab, ETH Zurich, Switzerland, use a digital genome design algorithm in conjunction with large-scale chemical DNA synthesis to physically produce artificial genomes and understand the code of life at the molecular level. The lab also uses systems and synthetic biology approaches to define essential genes across species that serve as the genetic parts to build microbial genomes for applications in sustainable chemistry, medicine and agriculture.
The research team has physically produced Caulobacter ethensis-2.0, the world's first fully computer-generated genome. Using a natural freshwater bacterium as a starting point, the researchers computed the ideal DNA sequence for chemical manufacturing and construction of a minimized genome solely composed of essential functions. In the design process, more than one-sixth of all of the 800,000 DNA letters in the artificial genome were replaced and the entire genome was produced as a large ring-shaped DNA molecule. While a living cell does not yet exist, gene functions have been tested across the entire genome design. In these experiments, researchers found that approximately 580 of the 680 artificial genes were functional, demonstrating the promise of the approach to produce designer genomes.
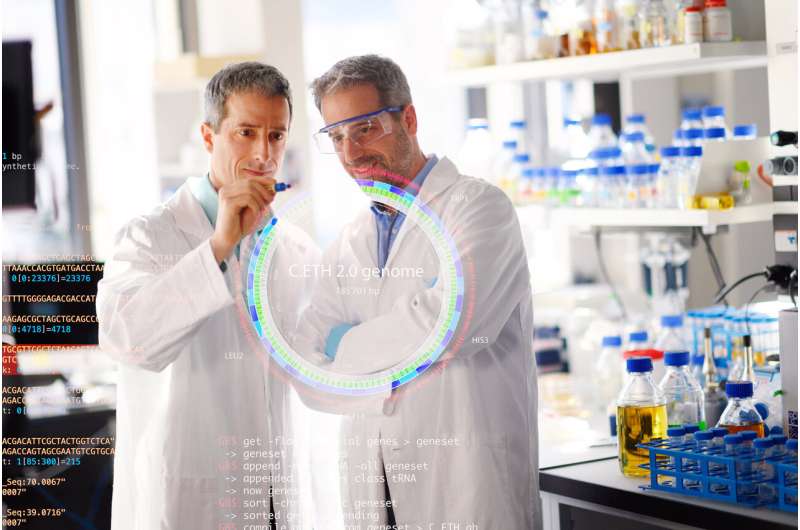

During the AAAS 2020 session, "Synthetic Biology: Digital Design of Living Systems" (February 14th, 2020), Christen will discuss possible future applications of synthetic genomes for industrial purposes and health benefits. He will also talk about the need for profound discussions in society about the challenges and purposes for which this technology can be used and, at the same time, about how potential for abuse can be prevented.
Professor Christen will be joined by Professor David Baker, University of Washington who will speak about Designer Proteins and Joyce Tait, University of Edinburgh, Edinburgh Scotland, who will speak on Risk Regulation, Uncertainty, and Ethics.
More information: Jonathan E. Venetz et al, Chemical synthesis rewriting of a bacterial genome to achieve design flexibility and biological functionality, Proceedings of the National Academy of Sciences (2019). DOI: 10.1073/pnas.1818259116
Computer-Generated Genomes AAAS 2020 session: aaas.confex.com/aaas/2020/meet … gapp.cgi/Paper/26042
Journal information: Proceedings of the National Academy of Sciences
Provided by ETH Zurich