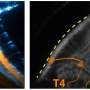
Surveying all the proteins on a neuron's surface
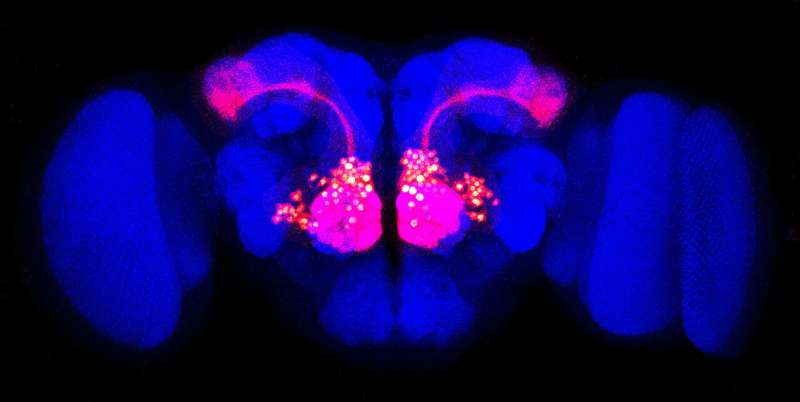
Scientists have found a new way to home in on the proteins covering a particular cell's surface. The feat offers insight into how brain cells form intricate networks during development.
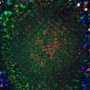
As if casting a tiny net, a new technique has rounded up all the proteins on the surface of neurons in the brains of fruit flies. The roundup uncovered 20 new molecules involved in wiring the developing brain.
The find furthers scientists' understanding of how neurons in the brain form complex networks, researchers report January 16, 2020, in the journal Cell. And it demonstrates for the first time that this protein-finding method actually works in intact brain tissues—not just cells grown in the lab, says study coauthor and Howard Hughes Medical Institute Investigator, Liqun Luo.
That's important because the tissue environment is crucial for cells' development, and lab cell cultures can't replicate it. Until now, scientists had no way to monitor all the proteins on cell surfaces in complex tissues like the brain. The new approach provides a way to survey this previously mysterious landscape.
"What really blew me away was the biological follow-up," says biochemist Matthias Mann of the Max Planck Institute of Biochemistry. Luo's team was able to find a trove of proteins whose biological role was previously unknown, says Mann, who was not involved with the work.
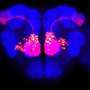
Cell surfaces are incredibly dynamic places, especially for cellular communication, says Luo, a neurobiologist at Stanford University. In the nervous system, proteins on the surfaces of nerve cells help the cells find each other and link up. Luo's team wanted a complete view of the proteins that direct connections in the developing fly brain. The researchers focused on proteins involved in forming olfactory networks, which control a fly's sense of smell.
Luo, along with his doctoral student Jiefu Li and collaborators at Stanford and the Broad Institute of MIT and Harvard, modified a method pioneered by study coauthor Alice Ting. In this method, called proximity labeling, researchers use an enzyme to add a molecular tag to a particular protein of interest, plus all the neighboring proteins. Researchers can then identify the tagged proteins using a chemical analysis called mass spectrometry.
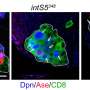
Luo's team added a new twist to the technique. They made the enzyme target proteins on fruit fly olfactory neurons at a particular point in brain development: when neurons are making decisions about which connections to form. The team compared the proteins present in adult cells with those present in the developing brain. "The difference is actually very striking," Luo says.
The team identified 20 proteins that were more abundant on the surfaces of developing neurons and knocked them down one by one to see if their absence had an effect on brain wiring. Surprising even to the researchers, all 20 were involved in wiring the fly olfactory network. What's more, many of the proteins they found hadn't even been known to play a role in neural development.
Luo and Li hope their approach will be useful for researchers in other fields as well. Li says it could be applied to immunology as well as to understanding how organs develop or modeling disease. For example, he adds, cell surface proteins are altered in cancer cells, so profiling those proteins could help scientists understand how cancer cells behave within tissues.
"I'd love to use [this technique]," says Joshua Sanes, a neuroscientist at Harvard University who was not involved in the research. Like Luo, Sanes (who is a member of HHMI's Scientific Review Board) is interested in how neurons form the precise patterns of connections that lead to all the complex neural circuitry underlying behavior. But he studies the brains of mammals, not flies. So, first, Sanes says, the method will have to be optimized for mammalian cells—a goal that has so far been elusive.
More information: Jiefu Li et al. Cell-Surface Proteomic Profiling in the Fly Brain Uncovers Wiring Regulators, Cell (2020). DOI: 10.1016/j.cell.2019.12.029
Journal information: Cell
Provided by Howard Hughes Medical Institute