Modified RNA has a direct effect on DNA

An article titled "m6A RNA modification as a new player in R-loop regulation," by the Dynamic Gene Regulation research group led by Arne Klungland at IMB, was published in the January edition of Nature Genetics.
Clarification: Modified RNA is distinct from messenger RNA, which simply comprises instructions for building proteins; messenger RNA, the basis of two prominent COVID vaccines, does not affect DNA.
Following a new collaboration between UiO and research groups in Nottingham and Oxford, it has now been revealed that RNA has a direct effect on DNA stability, according to Professor Klungland's research.
He believes the discovery will provide the health service with an important tool, since many studies have shown that the regulation of modifications to RNA is important for the development of cancer.
If genes that are important for the chemical compound 6-methyladenine are completely removed, this results in neurodegeneration in both mice and humans.
Where and how
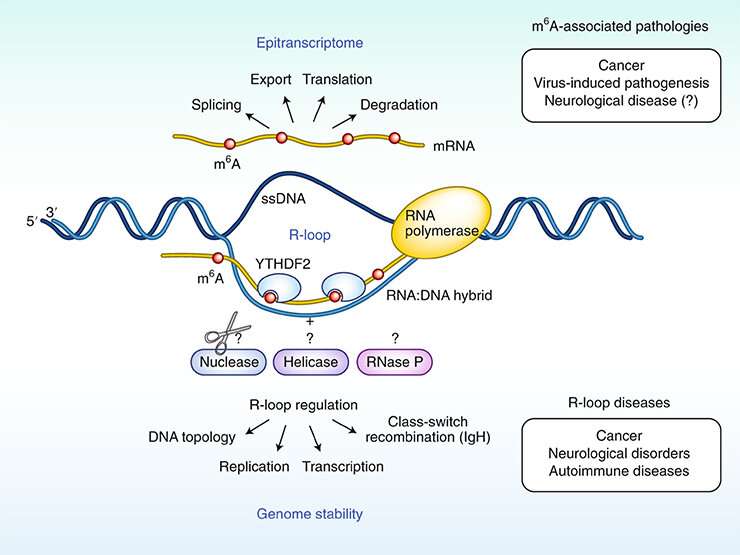
In areas of DNA where RNA binds to one of the DNA threads in such a way that the complementary DNA thread becomes the sole thread (R-loop structures), the DNA stability will change if RNA is chemically modified by m6A.
"Several research groups are now working together to study what effect this can have on the DNA molecule. We already know that R-loop areas are associated with sequences of DNA containing active genes and that this can lead to chromosomal breakage and the loss of genetic information," explains Klungland.
New field of research
Normally, epigenetic gene regulation is studied by examining dynamic modifications of DNA and proteins—so-called epigenetic modifications. The modifications can turn genes on or off without changing the underlying genetic code.
Less than 10 years ago, it was discovered that dynamic modifications also exist in RNA and that these have an important role to play in gene regulation
Important modification
The most common modification is on mRNA is 6-metyladenin (m6A). It has now been shown that this modification is essential for the survival of cells and model (non-human) organisms.
Over the last five years, there has been an enormous increase in the amount of research into RNA modifications—a field called epitranscriptomics.
One of the first studies in this field of research was the result of a collaboration between research groups in Chicago, Beijing and Oslo (Zheng, Dahl et al., Molecular Cell, 2012, 49, 18-29).
More information: Aline Marnef et al. m6A RNA modification as a new player in R-loop regulation, Nature Genetics (2019). DOI: 10.1038/s41588-019-0563-z
Journal information: Nature Genetics
Provided by University of Oslo