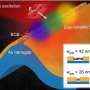
Pathways of disease spillover among domestic and wild sheep and goats in the western United States

A new large-scale genetic study has determined that domestic sheep and goats are the source of bronchopneumonia in bighorn sheep and mountain goats in the western United States, according to a research team led by Pauline Kamath, University of Maine assistant professor of animal health.
Using nearly 600 isolates collected over a 33-year period, the five-member team studied the genetic structure of the bacterium Mycoplasma ovipneumoniae, the primary causative agent of bronchopneumonia, in the domestic and wild sheep and goats to better understand transmission and spillover dynamics of the pathogen.
Spillover diseases have significant consequences for human and animal health, including wildlife conservation efforts, according to the researchers, writing in Scientific Reports, a Nature Research journal. In particular, bronchopneumonia has contributed to historical declines of bighorn sheep across western North America.
The disease is believed to have originated following exposure to domestic sheep introduced by European settlers and continues to impede conservation efforts to reestablish bighorn sheep across the species' range.
The genetic data in the new study identify domestic sheep as the primary infection reservoir, and source of M. ovipneumoniae spillover to bighorn sheep and mountain goats. Domestic goat strains were genetically distinct, but were also found to spill over into bighorn sheep. Following spillover events, the pathogen may persist in wildlife populations for extended periods.
The researchers found a high number of bacterial strains in domestic sheep flocks, the majority of which (77%) harbored unique strains. In domestic goats, there also was a large proportion of herds with unique strains (46%).
In contrast, 9% of bighorn sheep herds were infected with unique strains of M. ovipneumoniae. One of the strains detected in bighorn sheep was shared with domestic sheep; another with domestic goats.
The data suggest that the ability to predict M. ovipneumoniae spillover into wildlife populations may remain a challenge given the high strain diversity in domestic hosts and need for more comprehensive pathogen surveillance, the researchers write. Knowledge of pathogen movement, invasion frequency and sources is key to helping predict the ability of spillover host species to persist and recover from pathogen infections.
More information: Pauline L. Kamath et al. Genetic structure of Mycoplasma ovipneumoniae informs pathogen spillover dynamics between domestic and wild Caprinae in the western United States, Scientific Reports (2019). DOI: 10.1038/s41598-019-51444-x
Journal information: Scientific Reports
Provided by University of Maine