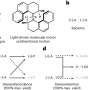
An improved method for protein crystal structure visualization

X-ray crystal structure visualization technique has been known for over a hundred years. While it keeps improving, it is extremely difficult to focus rays on objects that are invisible to the naked eye, such as proteins. However, to get a clear image and effectively visualize the structure of a crystal, a sample should be positioned correctly. An international team of scientists suggested an optic system to help see a protein crystal in X-rays and place it in the center of a beam. The results of the study were published in the Structural Biology journal.
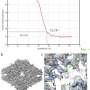
During crystallization atoms are arranged in a 3-D lattice structured in a specific way. The distances between the atoms in that lattice are determined by the atoms themselves. The X-ray wavelength is comparable to interatomic distances, so the rays can be refracted on the planes. Due to this effect one can analyze crystal structure. The X-ray images show the distances between the planes. Based on this information it is possible to determine what atoms are in the lattice and how they interact with each other. In proteins studies, for example in the search for new drugs, their structure can be determined on the level of basic atomic groups (amino acids).
The main issue of X-ray crystallography is that microscopic protein crystals are very difficult to position in the center of an X-ray beam, and hence that X-ray diffraction image may be blurry. Moreover, if the exact position of a crystal is unknown, one has to scan the whole sample. This increases the time of exposure to very intensive X-rays. Biological molecules start to denature under that exposure.
An international team of scientists developed an optic system allowing one to see a sample in X-rays and discern its position and orientation relative to the beam. Just like with a regular optical microscope it can move the sample, adjust ray intensity, and focus the beam. Such a system can significantly reduce the time of analysis and thus preserve the integrity of the molecules. Scientists have demonstrated how the system works on the example of a crystal of the antibacterial protein lysozyme. The quality of X-ray diffraction images turned out to be much higher after the positioning of the sample in the X-ray beam.
"Our system is now successfully used in the international research center by the DESY synchrotron in Hamburg, where the laboratories of the world's leading universities carry out their crystal structure studies. In the future, we plan to automate the crystal positioning process using neural networks," said Prof. Anatoly Snigirev, the head of the Science and Research Center "Coherent X-Ray Optics for Megascience Installations," at Kant Baltic Federal University.
More information: Maxim Polikarpov et al, Visualization of protein crystals by high-energy phase-contrast X-ray imaging, Acta Crystallographica Section D Structural Biology (2019). DOI: 10.1107/S2059798319011379
Provided by Immanuel Kant Baltic Federal University