Credit: CC0 Public Domain
Bio-Protection Research Centre scientists and collaborators have made a discovery that potentially opens the door to new medicines and biological pesticides.
In work that began with a Ph.D. thesis, and continued through postdoctoral work, they have shown that scientists have been missing potential new drugs and biopesticides, dismissing the genes that make them as non-functional variants.
"This is a fundamental discovery," says Prof Barry Scott, who is based at Massey University. Prof Scott mentored Daniel Berry, the scientist who started investigating the "junk genes" as part of his Ph.D. research in 2012.
Now Dr. Berry is the lead author on the paper outlining the discovery, which has just been published in the highly prestigious journal Proceedings of the National Academy of Sciences.
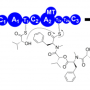
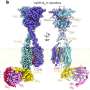
Dr. Berry's discovery involves a group of "modular enzymes", which naturally occur in fungi. Modular enzymes are molecular assembly lines that make bioactive substances, which scientists have harnessed to work as medicines, or as natural pesticides or growth enhancers in agriculture and horticulture.
The antibiotic penicillin is perhaps the best-known product of a modular enzyme.
In 2012, while researching for his Ph.D. thesis, Dr. Berry was investigating synthesis of peramine, a natural pasture pest-repellent made by a modular enzyme of the fungus Epichloë.
"A lot of the genes had been mucked up and didn't make peramine," says Prof Scott. "But one of those gene variants kept occurring, so although it looked like it wasn't functioning, Daniel realised it had to serve some purpose, and decided to investigate."
Prof Scott said for some time their investigations led nowhere. "That's often the nature of science—it can take a long time."
The breakthrough came when Daniel went to Germany to work with Dr. Helge Bode, an expert in modular enzymes. With Dr. Bode's help, he discovered the modular enzyme from the variant gene produced a product they named cycloProArg, which is likely to be bioactive.
"It was an exciting day when we were finally able to identify what this gene was making," says Dr. Berry. "This discovery really opened the door for us, as it enabled identification of an additional three products that are made by other variants of this gene."
"We don't yet know precisely what cycloProArg does—that will be the subject of further research," says Prof Scott. "But we know that as well as providing protection from pests, Epichloë promotes pasture growth and has lots of other beneficial effects. CycloProArg is probably responsible for one of those benefits."
The real breakthrough, he says, is the discovery that modular enzymes make more than one bioactive product.
"Previously we thought one enzyme, one product," says Prof Scott. "But we now know this is wrong.
"This fundamentally changes what we know, and what we should be looking for.
"Scientists in many fields—including medicine—should go back and take a look at previously discarded 'junk genes', because some of these may actually produce useful bioactive products."
More information: Daniel Berry et al. Efficient nonenzymatic cyclization and domain shuffling drive pyrrolopyrazine diversity from truncated variants of a fungal NRPS, Proceedings of the National Academy of Sciences (2019). DOI: 10.1073/pnas.1913080116
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Otago