Researchers show how opportunistic bacterium defeats competitors
Researchers at the University of São Paulo (USP) in Brazil have described a system present in a species of opportunistic bacterium found in hospital environments that injects a cocktail of toxins into competing bacteria and completely eliminates them. This discovery can be used in the future development of new antimicrobial compounds.
The study was supported by São Paulo Research Foundation—FAPESP and published in PLOS Pathogens by researchers affiliated with USP's Chemistry Institute (IQ) and Biomedical Sciences Institute (ICB).
The researchers discovered that Stenotrophomonas maltophilia uses a secretion system that produces a cocktail of toxins and injects them into other microorganisms with which it competes for space and food. In addition, the researchers characterized one of 12 proteins in the cocktail—Smlt3024—and observed that this molecule alone can considerably reduce the rate at which other bacteria replicate.
"We believe these toxins can be explored as a form of treatment in the future. Just as antibiotics can come from other bacteria, we are exploring this arsenal used by bacteria themselves to kill other species of pathogen," said Ethel Bayer-Santos, a researcher at ICB-USP.
The study was part of her postdoctoral project at IQ-USP, which was also supported by FAPESP. She currently has a Young Investigator grant from FAPESP.
Bacterial secretion systems are protein complexes present on the cell membranes of bacteria for the secretion of substances. These systems are used by pathogenic bacteria to secrete virulence factors and invade host cells.
"The Type IV secretion system [T4SS], as it is known, secretes proteins and DNA into other cells or the extracellular medium. We recently showed that it is present in Xanthomonas citri, the pathogen that causes citrus canker. We have now found it in this bacterium [S. maltophilia], which is frequently isolated from water and soil but can become an opportunistic pathogen in hospital environments," said Shaker Chuck Farah, a professor in IQ-USP and principal investigator for the study.
The research is linked to two Thematic Projects funded by FAPESP: "Cyclic di-GMP signaling and the Type IV macromolecule secretion system in Xanthomonas citri" and "Structure and function of bacterial secretion systems".
The group's discovery of the secretion system in X. citri was reported in a paper published in Nature Communications. Another article by the same group, published in 2018 in Nature Microbiology, describes the atomic structure of much of the secretion system using cryogenic electron microscopy (cryo-EM), an imaging technique that enables scientists to observe the 3-D structure of biomolecules as they move and interact.
Bacterial warfare
Competition between microorganisms determines which species will dominate or be eradicated from a specific habitat. Bacteria have several mechanisms that can reduce the multiplication of competitors or even kill them. One is T4SS, which itself consists of more than 100 components originating from 12 different proteins present on the surface of their cells.
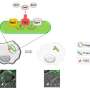
The researchers found that the secretion systems in X. citri and S. maltophilia contain antidotes to the toxins they secrete. These antidotes are required to protect the microorganisms from their own weapons. On approaching another species of bacterium, the attackers inject the toxins into the competitor by mere contact or using a pilus, a microscopic filament that functions like a harpoon.
To test the action of the system, the researchers performed several experiments involving competition between bacteria. In one experiment, they placed specimens of S. maltophilia and Escherichia coli, a species widely used in the laboratory, in the same medium. E. coli had barely begun to multiply when S. maltophilia approached and eliminated its rivals. The same effect was observed against Klebsiella pneumoniae and Salmonella typhi, as well as Pseudomonas aeruginosa, a bacterium that infects cystic fibrosis patients.
Another experiment consisted of testing the effect of Smlt3024 alone. Specimens of E. coli that expressed the toxin in the cytoplasm exhibited no growth alterations. However, when, in addition to the gene for Smlt3024, they also had a signal sequence—a set of certain amino acids that changes the location of the protein—the toxin was directed to the periplasm (the space between the inner and outer membranes in bacteria), exactly as the original system does in S. maltophilia and X. citri. In this case, cell division in E. coli was considerably reduced, demonstrating the effect of the protein.
Although both S. maltophilia and X. citri have the same system, they can kill each other using T4SSs, as another experiment showed. Experiments also showed that key components of one system can be replaced by homologues from another and that the T4SS in one bacterium can secrete the toxins of another.
All this is evidence of similarities in the structures that can be studied in the future to explore similar T4SSs (homologues) identified in tens of other bacterial species.New studies may allow other researchers to develop molecules that inhibit the action of this system for use in the discovery of new drugs.
"We can screen drugs capable of binding to this system and inhibiting some important stages in the transfer of toxins from one cell to another. This could reduce the survival of these bacteria in some environments," Farah said.
"The toxin itself could be used as an antimicrobial agent, but other studies need to be done to test this possibility," Bayer-Santos said.
The study also has three other authors supported by FAPESP, and all are affiliated with IQ-USP: Bruno Yasui Matsuyama, who has a postdoctoral scholarship from FAPESP; Gabriel Umaji Oka, who also has a postdoctoral scholarship from FAPESP; and William Cenens, whose postdoctoral scholarship ended in May 2019.
More information: Ethel Bayer-Santos et al. The opportunistic pathogen Stenotrophomonas maltophilia utilizes a type IV secretion system for interbacterial killing, PLOS Pathogens (2019). DOI: 10.1371/journal.ppat.1007651
Diorge P. Souza et al. Bacterial killing via a type IV secretion system, Nature Communications (2015). DOI: 10.1038/ncomms7453
Germán G. Sgro et al. Cryo-EM structure of the bacteria-killing type IV secretion system core complex from Xanthomonas citri, Nature Microbiology (2018). DOI: 10.1038/s41564-018-0262-z
Journal information: PLoS Pathogens , Nature Communications , Nature Microbiology
Provided by FAPESP