Coordinating the removal of RNA-DNA hybrids
Two research teams led by Professors Brian Luke and Helle Ulrich at the Institute of Molecular Biology have deciphered how two enzymes, RNase H2 and RNase H1, are coordinated to remove RNA-DNA hybrid structures from chromosomes. RNA-DNA hybrids are important for promoting normal cell activities like gene regulation and DNA repair, but having too many is also a risk for DNA damage and can lead to neurodegenerative disease and cancer.
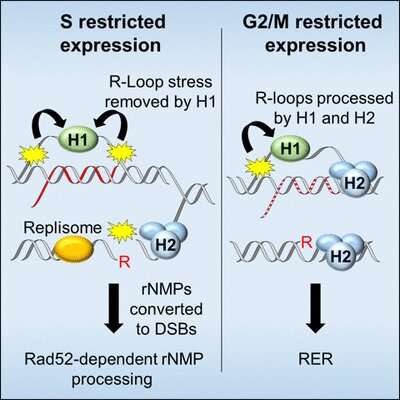
In their article, which was published today in Cell Reports, Luke and Ulrich show that the enzyme RNase H2 removes RNA-DNA hybrids primarily after DNA replication. Any remaining RNA-DNA structures are then removed by RNase H1, which acts independently of cell cycling.
DNA is normally found as a stable, double-stranded structure. However, DNA also sometimes interacts with RNA to form RNA-DNA hybrid structures that regulate gene expression and DNA repair. R-loops are a special type of RNA-DNA hybrid in which an RNA strand binds to one strand of a DNA molecule and pushes out the other DNA strand so that it is exposed as a single-stranded loop. R-loops can regulate gene activity, but also quickly become dangerous because incorrect removal can damage the DNA, potentially causing mutations. Therefore, excess R-loop formation can be toxic for cells—indeed, mutations in R-loop removal proteins are known to contribute to neuroinflammatory diseases and cancer.
R-loop removal is catalysed by the enzymes RNase H1 and RNase H2, which degrade the RNA strand. In addition, RNase H2 also has a secondary ability to excise single ribonucleotides, which can sometimes be mistakenly incorporated into DNA by polymerases in a process known as ribonucleotide excision repair (RER). Previous studies showed that mutation of RNase H2 disrupted genome stability more than RNase H1 mutation, suggesting that RNase H2 has a more important role in maintaining genome stability. However, it was never fully understood how these important enzymes are coordinated.
To dissect the distinct roles of RNase H1 and H2 in R-loop removal, the Luke lab engineered yeast to express RNase H1 and H2 only during specific phases of the cell cycle and then exposed them to methyl methanesulfonate (MMS), an agent that increases R-loop formation. Only yeast that could effectively remove R-loops would survive, while those with impaired R-loop removal would not survive.
With support from the Ulrich lab, they found that yeast expressing RNase H2 exclusively during G2 (the 'growth phase' of the cell cycle following DNA replication) were resistant to MMS, whereas yeast expressing RNase H2 only during S phase (the DNA replication phase) were more sensitive. This suggested that RNase H2 primarily acts to process R-loops during G2. In contrast, yeast expressing RNase H1 in either G2 or S phase were both able to survive in MMS. Surprisingly, RNase H2 expression in S phase actually induced more DNA damage, which required a special type of DNA repair called homologous recombination to fix. This pathway was not previously known to act during S phase. Therefore, this study may have revealed an unexplored repair pathway which counters damage caused by RNase H2 activity during DNA replication.
These results may explain why cells have evolved two different RNAse H enzymes. Luke clarifies: "We think that RNase H2 is the 'housekeeping' enzyme that repairs the majority of RNA-DNA hybrids, but it is strictly regulated by cell cycling and acts only in G2 phase, or after DNA replication." Luke and Ulrich speculate this may be because the additional RER activity of RNase H2 creates nicks in the DNA, which are a risk for double-strand breaks during DNA replication in S phase. Therefore, cells may have also evolved a secondary enzyme, RNase H1, which does not have RER activity and can act in all phases of the cell cycle, including S phase.
These findings help us to further understand how cells repair DNA damage associated with RNA-DNA hybrids and how impairment of this process contributes to disease.
More information: Arianna Lockhart et al, RNase H1 and H2 Are Differentially Regulated to Process RNA-DNA Hybrids, Cell Reports (2019). DOI: 10.1016/j.celrep.2019.10.108
Journal information: Cell Reports
Provided by Universitaet Mainz