New proteomics technique gives insights into ubiquitin signalling
Australian researchers are among the first in the world to have access to a new approach to understand intricate changes that control how proteins function in our cells in health and disease.
The new proteomics technique called 'ubiquitin clipping' allows researchers to create high-definition maps of how proteins are modified by a process called ubiquitination. The technique provides a new level of detail for understanding the role of ubiquitination in cells, and could uncover subtle changes that contribute to a range of diseases including cancer, inflammatory conditions and neurodegenerative disorders.
The research, which was published today in Nature, was led by Walter and Eliza Hall Institute researcher Professor David Komander, who undertook the work at the MRC Laboratory of Molecular Biology in Cambridge, UK.
At a glance
- A new proteomics technique has been developed to study protein ubiquitination, modifications of proteins in cells that can impact their function.
- The technique, called 'ubiquitin clipping', enables researchers to map protein ubiquitination in unprecedented detail.
- Ubiquitin clipping is now established at the Walter and Eliza Hall Institute, enabling Australian scientists to investigate new aspects of ubiquitin signalling, including its links to a range of diseases.
Ubiquitin architecture
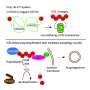
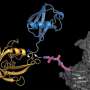
Ubiquitin is a small protein that can link to other proteins in a cell, either as a single unit or in longer straight or branched chains. Professor David Komander, who heads the Institute's recently established Ubiquitin Signalling division, said protein ubiquitination could impact all cellular processes.
"Ubiquitination can change how proteins function, potentially altering their activity, redirecting them to different parts of the cell, or regulating their interactions with other proteins. One of the best-known examples of ubiquitination is when it targets specific proteins for destruction, regulating the levels of the protein in the cell, but we now know there are many more subtle and complex roles for ubiquitin signalling," he said.
"The 'architecture' of ubiquitin chains can be complex with many branches that influence its impact on proteins, yet until now it has been almost impossible for researchers to detect and distinguish between different branching structures. This has limited the experiments that were possible to understand the role of ubiquitination in disease processes."
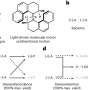
The new 'ubiquitin clipping' technique, which was developed by Professor Komander and his colleagues at the University of Cambridge and the University of Vienna, enables scientists to measure different ubiquitin chain architectures by pretreating protein samples, and then analysing them using electrophoresis and mass spectrometry.
"Ubiquitin clipping has enabled us to reveal a whole new level of complexity in ubiquitin signalling. In our pilot experiments, we discovered branched ubiquitin chains are much more common than previously thought. We could also study combinations of modifications on ubiquitin and other proteins—a feat that was until now rather difficult," Professor Komander said.
"This is a revolutionary technique that simplifies ubiquitin research, enabling a new level of detailed experimentation. It's the difference between describing a house based solely on the number of walls, windows and doors it has, versus looking at the detailed architectural plans."
New insights into diseases
Altered ubiquitination of proteins has been implicated in a range of diseases, including cancer, inflammatory diseases and neurodegenerative disorders such as Parkinson's disease. Professor Komander said the ubiquitin clipping technique was already being applied to study patient samples.
"My Walter and Eliza Hall Institute colleague Dr. Rebecca Feltham is using ubiquitin clipping to look for protein ubiquitination patterns in samples from patients with rheumatoid arthritis, a complex inflammatory disease. This could give new insights into how this disease develops and responds to existing therapies," he said.
"Ubiquitination is also a promising target for the development of new drugs. Ubiquitin clipping will be a critical aspect of my team's drug discovery research.
"It's exciting that Australian researchers are among the first in the world to have access to ubiquitin clipping, and I'm looking forward to seeing the technique underpin many exciting discoveries, including through our collaborations with other research groups."
More information: Insights into ubiquitin chain architecture using Ub-clipping, Nature (2019). DOI: 10.1038/s41586-019-1482-y , nature.com/articles/s41586-019-1482-y
Journal information: Nature
Provided by Walter and Eliza Hall Institute