Scientists teach old worms new tricks
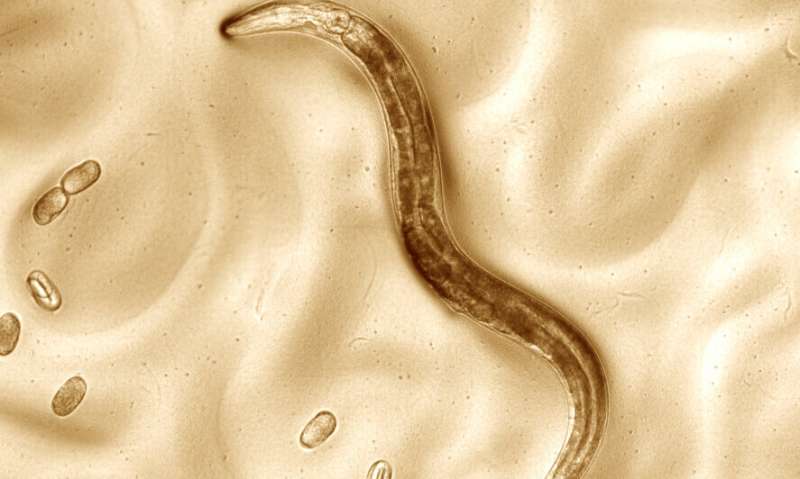
Model organisms such as yeast, fruit flies, and worms have advanced the study of genomics, eukaryotic biology, and evolution. An important resource for any model organism is a near-complete reference genome from which a multitude of scientific questions can be answered. Caenorhabditis elegans have been widely studied due to their short generation time and transparent anatomy and were one of the first multicellular organisms sequenced, yet gaps in their reference genome remain.
Three studies, published today in Genome Research, provide novel insights into C. elegans genomics and gene expression. Advances in sequencing technologies toward longer reads have facilitated genome assembly, finishing, and the sequencing of highly repetitive regions. In a study by Yoshimura and colleagues, researchers used a combination of short- and long-read assemblies to generate a more complete reference genome of a modern laboratory strain of C. elegans. The new sequence has an extra two million nucleotides that were absent from the previous sequence, which include highly repetitive regions and approximately 50 new genes. Likewise, Kim and colleagues used long-read sequencing to construct a high-quality reference genome of a wild C. elegans strain for comparative studies, detecting new regions at chromosomes ends that maintain genome integrity and more than 600 genes divergent between the wild strain and a laboratory strain.
Knowledge of where and when these genes are expressed is then key to understanding how organism developed and how tissues carry out their specific functions. Robert Waterston and colleagues utilized C. elegans to study spatial and temporal gene expression in a cell- and tissue-specific manner during development. This gene regulatory network library provides a framework for studying cell lineage from egg to adulthood.
These studies expand the usefulness of this model organism and provide an important resource to C. elegans biologists. Equally, the development of advanced techniques may facilitate the generation of more complete genome assemblies and the discovery of context- and tissue-specific gene regulation where small cell niches may play important biological roles in development, immune response, and cancer.
More information: Yoshimura J, Ichikawa K, Shoura M, Artiles K, Gabdank I, Wahba L, Smith C, Edgley M, Rougvie A, Fire A, Morishita S, and Schwarz E. 2019. Recompleting the Caenorhabditis elegans genome Genome Res doi: 10.1101/gr.244830.118.
Kim C, Kim J, Kim S, Cook D, Evans K, Andersen E, Lee J. 2019. Long-read sequencing reveals intra-species tolerance of substantial structural variations and new subtelomere formation in C. elegans Genome Res doi: 10.1101/gr.246082.118
Warner A, Gevirtzman L, Hillier L, Ewing B, Waterston R. 2019. The C. elegans embryonic transcriptome with tissue, time, and alternative splicing resolution Genome Res doi: 10.1101/gr.243394.118
Journal information: Genome Research
Provided by Cold Spring Harbor Laboratory Press