LDAIR, a lncRNA regulates seasonal changes in stress response
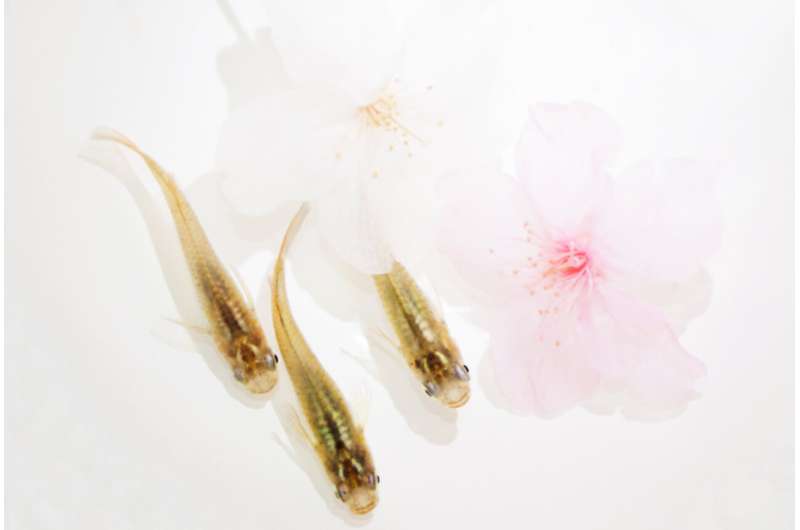

Biologists at the National Institute for Basic Biology and the Institute of Transformative Bio-Molecules (ITbM) at Nagoya University have discovered that long non-coding RNA regulates seasonal changes in stress response in medaka fish. The results of this study were reported in Nature Ecology & Evolution.
In temperate zones, the environment fluctuates greatly depending on the season, and animals living in these areas adapt to these changing environments. A research group from the National Institute for Basic Biology and Nagoya University in Japan uncovered the underlying mechanism of seasonal changes in stress response.
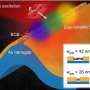
Although the Greek philosopher Aristotle described seasonal changes in animal behavior in his "Historia Animalium (History of Animals)", their underlying mechanisms remain unclear. During breeding seasons, animals show stress responses, such as self-protective and escape behaviors when confronted with such dangers as predation, interspecific competition and harsh weather. These behaviors are critical for animals to survive in changing environments. In order to investigate seasonal changes in stress responses, the research group examined medaka, a small fish inhabiting rice fields and streams. Contributing researcher and Ph.D. student Tomoya Nakayama said, "We examined genome-wide gene expression analysis during the transition from short-day to long-day conditions, and identified photoperiodically regulated genes."
Among the photoperiodically regulated genes, the researchers focused on a novel, long non-coding RNA (lncRNA), LDAIR (long-day-induced antisense intronic RNA), whose robust rhythmic expression was induced only under long-day conditions. lncRNA are defined as transcripts longer than 200 nucleotides that are not translated into protein. Although recent studies found that there are as many lncRNA strands as protein-coding genes in the human genome, the functional significance of most lncRNA remains unknown.
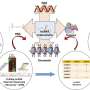
Using a genome-editing technique, the research group generated LDAIR knockout medaka to understand its function. As a result, it became clear that LDAIR regulates a gene neighborhood including corticotropin-releasing hormone receptor 2 (CRHR2). Corticotropin-releasing hormone (CRH) and its receptors, CRHR1 and CRHR2, are known to be involved in stress responses. Furthermore, upon conducting behavioral analysis, it was confirmed that LDAIR affects stress-associated self-protective behaviors.
Professor Yoshimura, the leader of the research group, said, "To cope with seasonal changes in the environment, animals often adapt their behavior. We found the involvement of lncRNA LDAIR in this adaptive behavior."
Recent studies concerning plants have also reported that photoperiodically regulated lncRNAs regulate photomorphogenesis, cotyledon greening, and flowering. Although the sequences of lncRNAs are not conserved, lncRNA appears to play an important role across species in the adaptation mechanism to seasonal changes in the environment.
More information: "Seasonal regulation of the lncRNA LDAIR modulates self-protective behaviors during the breeding season" Nature Ecology & Evolution (2019). DOI: 10.1038/s41559-019-0866-6
Journal information: Nature Ecology & Evolution
Provided by National Institutes of Natural Sciences