Scientists shine light on minute peptide changes affecting immune system
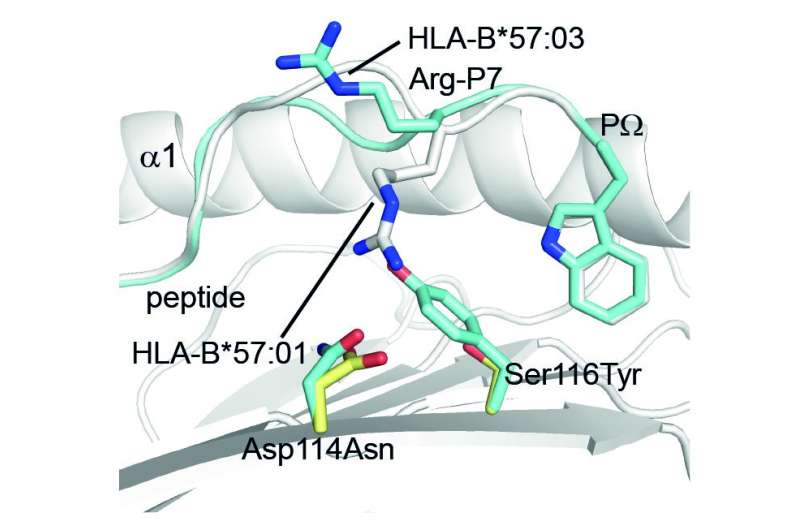
Human leukocyte antigen (HLA) molecules are a group of proteins that help the immune system identify foreign invaders such as viruses and bacteria and which impact a wide variety of clinical outcomes including infection and cancer. Now Monash Biomedicine Discovery Institute (BDI) researchers have made some important insights into the link between genetic diversity and the intricate changes in peptides bound to HLA molecules that influence these outcomes.
The genes that encode HLA molecules are extremely diverse—the most polymorphic part of the human genome—with different HLA variants displaying different peptide antigens at the cell surface. This genetic variation influences outcomes as wide-ranging as adverse drug reactions and whether a patient will react negatively to a donor transplant.
An international team of collaborators, led by Monash BDI's Professor Anthony Purcell, demonstrated that even very minor changes in genetic sequences in HLA molecules can impact dramatically on immune function.
Their findings, published today in Nature Communications, have important implications for understanding how the immune system recognises infected cells in different individuals.
The multidisciplinary team of researchers explored the impact of micropolymorphism on three closely related HLA molecules associated in different ways with drug hypersensitivity and HIV 'elite control' - where an individual's immune system seems to help keep HIV infection at bay.
"The only difference between these HLA molecules is a few amino acids strategically placed in the clefts of these molecules where peptide antigens sit," Professor Purcell said.
"We knew there might be subtle differences in the sort of peptides bound to these molecules, but what we wanted to do in a systematic way was to understand what the differences are and what sort of functional impact the differences might have," he said.
"It's quite easy now to map polymorphism, and map changes in amino acid sequences; the difficulty is really translating that to say what that might mean, to be able to predict to some extent how different alleles will select different peptide antigens and how this impacts on immunity."
"It's striking the way that with such minor differences in the HLA you are getting both differences in the peptide interaction with the HLA molecule and also in the shape of the peptides, which makes it a drastically different target for the immune system," Dr. Patricia Illing, joint first author, said.
Professor Purcell said the study, part of research spanning 25 years, provided fundamental insights that may help explain such processes as elite control in HIV or why individuals who have closely related HLA molecules can have problematic or 'taboo' mismatches in transplantation.
The knowledge gained may also help in the development of personalised vaccines.
"In my view, understanding how polymorphism impacts on antigen presentation or peptide presentation will actually aid the predictive tools that will be used to formulate vaccines, for example in cancer or infectious disease," he said.
It also has broader implications.
"The study helps us to understand how the immune system is evolving to try different ways of presenting peptides, and how the genetic diversity of the human population has been shaped by pathogens," he said.
This is the second study published recently in which Professor Purcell and other researchers have made insights into peptides; another paper published in Science Immunology on 'trans-spliced peptides', highlighted the unanticipated diversity of the immuno-peptidome.
More information: Patricia T. Illing et al, HLA-B57 micropolymorphism defines the sequence and conformational breadth of the immunopeptidome, Nature Communications (2018). DOI: 10.1038/s41467-018-07109-w
Journal information: Nature Communications , Science Immunology
Provided by Monash University