November 5, 2018 feature
Ready for its close-up—a bacterium's electron transport pathway
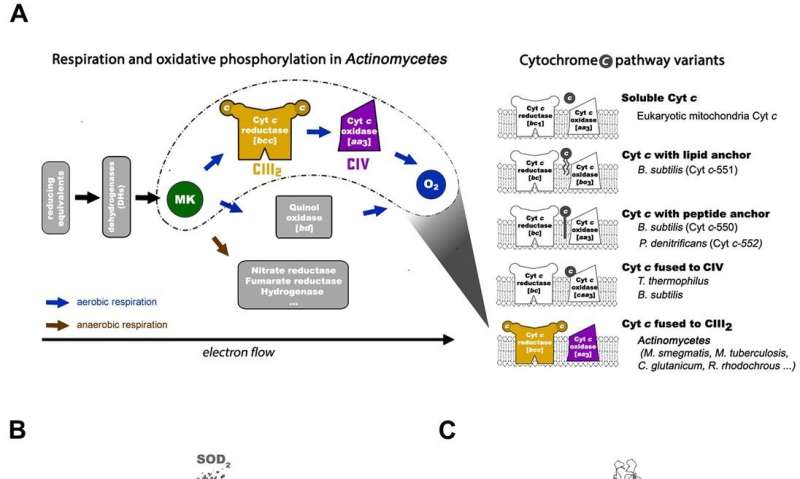
In a recent study conducted by Hongri Gong and colleagues, a respiratory supercomplex was isolated from the bacterium Mycobacterium smegmatis, and its structure was visualized at a resolution of 3.5 Å using cryo-electron microscopy (cryo-EM). The bacterium is a close relative to M. tuberculosis and a popular model used to study many other bacterial species. The detailed structure revealed how electrons were transferred in the cell in a process hitherto unseen.
Typically, chemical energy to synthesize adenosine triphosphate (ATP) and power cellular reactions is extracted during cellular respiration by coupling the oxidation of an energy source (sugars, fatty acids and amino acids) and the reduction of an electron acceptor (oxygen, sulfur, nitrate and sulfate). In aerobic cellular respiration, energy is extracted from electron donors to the terminal acceptor, oxygen, via the electron transport chain (ETC) to create a transmembrane proton gradient known as a proton motive force (PMF) that drives ATP synthesis. The new results now published in Science reveal a direct link for electron transfer between enzymes to represent a new mode of respiratory chain catalysis.
Quinones and cytochromes are two types of electron carriers in ETCs used to shuttle electrons to and from large macromolecular structures embedded in the membrane. Four membrane oxidoreductases are involved in the mitochondrial respiratory chain for electron transfer. These include complex I (NADH:ubiquinone oxidoreductase, CI), complex II (succinate:ubiquinone oxidoreductase, CII), complex III (bc1-type ubiquinol:cytochrome c oxidoreductase, bc1-type CIII) and complex IV (aa3-type cytochrome c oxidase, aa3-type CIV). By function, CIII can oxidize ubiquinol to ubiquinone and pass the electrons to soluble cytochrome c. Electrons are then shuttled to CIV, where oxygen is reduced to water. The transmembrane PMF is generated by proton pumping in CI, CIII and CIV.
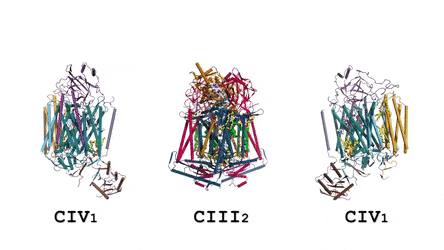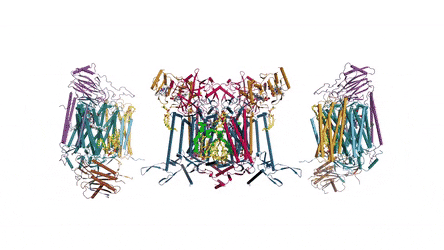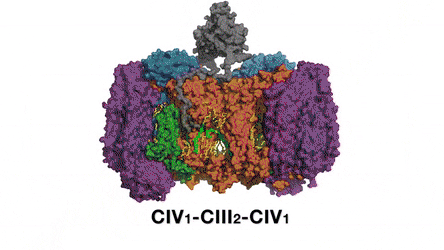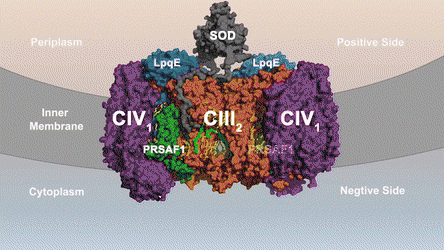
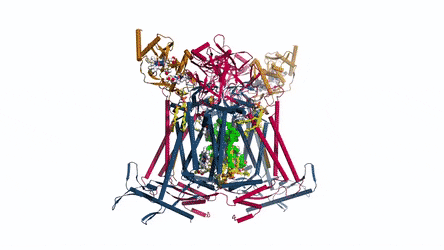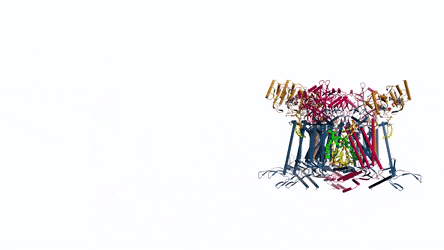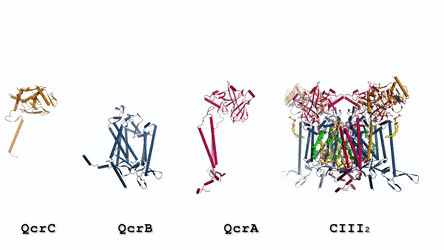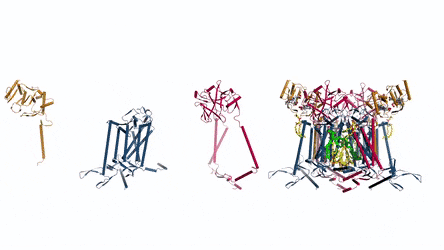
In the prokaryotic respiratory chain, the situation is more complicated. A complete pathway of electron flow has not yet been determined in the cell type due to its complexity. It is therefore necessary to understand the complete structure of a "supercomplex" involved during bacterial electron transfer to assist the goal. In the study, the researchers extracted and purified the complex from M. smegmatis to visualize the architecture using cryo-electron microscopy (cryo-EM) at a resolution of 3.5 Å. The structure provided crucial insights into the mechanism of direct electron transfer within a respiratory supercomplex. The dimensions of the supercomplex were in the range of 200 x 70 x 120 Å, in a symmetrized linear architecture completely different from previously reported respiratory supercomplexes. By composition, the linear dimeric CIV1-CIII2-CIV1 was arranged such that individual CIVs flanked the central CIII dimer on either side. The information revealed a direct link between enzymes during electron transfer, representing a new mode of respiratory chain catalysis. The detailed structural findings have potential to assist with antimycobacterial drug discovery efforts.
During bacterial cell culture experiments, the authors used an M. tuberculosis-like hydrogen peroxide-resistant M. smegmatis mutant strain. The cells were cultured and the membrane isolated as previously described. After cell culture, harvesting and cell lysis, cell membrane pellets were harvested to extract respiratory supercomplexes. The supercomplexes were characterized using optical spectroscopy, mass spectroscopy and 3,3'-diaminobenzidine (DAB) staining. To identify heme groups, selected fractions were analyzed by recording spectra before and after reduction with dithionate as described previously. The purified samples were analyzed using native mass spectroscopy to investigate the architecture and the individual structural components were analyzed using previously established protocols.
During cryo-EM analysis, the researchers used uranyl acetate (1 percent w/v) for negative staining, using 5 µl of the supercomplex sample at a concentration of 0.05 mg/ml, images were taken on a FEI Tecnai Spirit microscope operating at 120 kV for initial model building. The acquired images were processed using a low-resolution reconstruction of the supercomplex from 53 micrographs of the negative stained sample. For complete reconstruction of the supercomplex, the authors manually selected 7,600 micrographs from 8,200 original micrographs during cryo-EM image processing. All figures in the study were created using PyMOL or UCSF chimera.
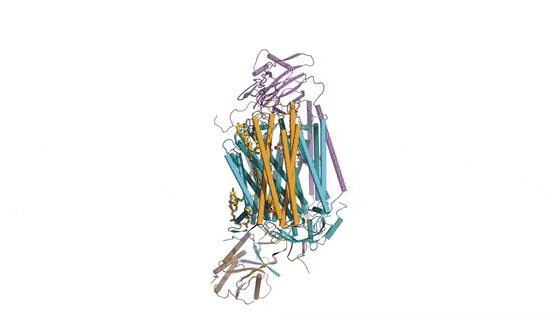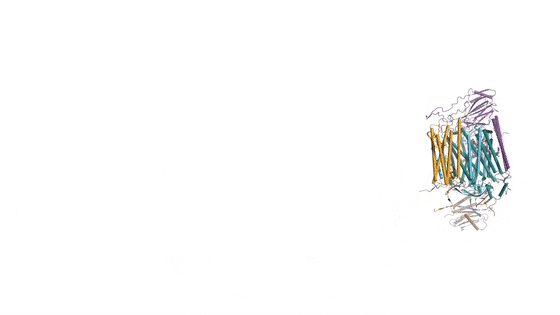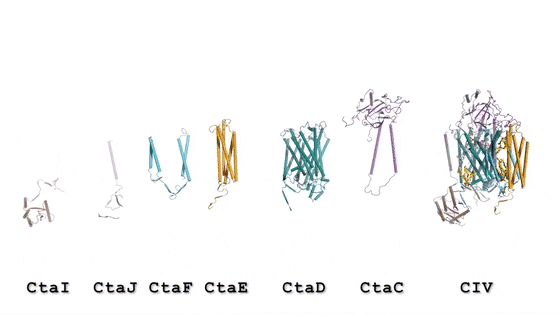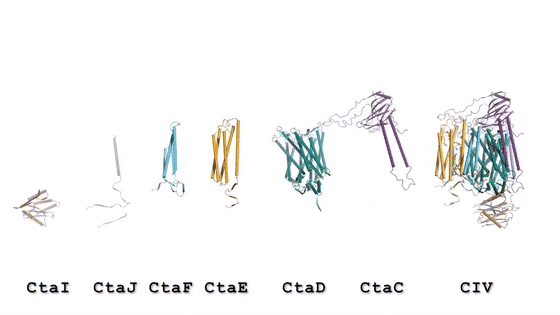
The authors revealed the cryo-EM structure of a CIII-CIV respiratory supercomplex of the M. smegmatis bacteria. The intra-complex electron transfer pathway ranged from quinol oxidation in CIII to oxygen reduction in CIV. The results showed a new mechanism for bifurcating electron transfer to ensure completion of the Q cycle (the net movement of protons across a lipid bilayer) for energy transduction. The association of a superoxide dismutase in the architecture of the system can protect against oxidative damage by reactive oxygen species (ROS). The architecture of quinone binding sites also provided a framework for future studies in structure-based antimicrobial drug discovery.
More information: Hongri Gong et al. An electron transfer path connects subunits of a mycobacterial respiratory supercomplex, Science (2018). DOI: 10.1126/science.aat8923
Eric F. Pettersen et al. UCSF Chimera?A visualization system for exploratory research and analysis, Journal of Computational Chemistry (2004). DOI: 10.1002/jcc.20084
Journal information: Science
© 2018 Science X Network