Better understanding of hydrogen peroxide regulation can lead to new insights into disease development
The team of prof. Joris Messens at the VIB-VUB Center for Structural Biology has provided new insights into the regulation of an important intracellular messenger molecule, hydrogen peroxide (H2O2), whose dysregulation has been linked to the development of several diseases, including cancer.
To fine-tune levels of H2O2, cells can sense changes in the concentration of H2O2 and respond by activating specific DNA regulation mechanisms. In bacteria, a protein called OxyR functions as such a H2O2-sensor. The exact mechanism of how OxyR senses H2O2 and changes its DNA binding properties, however, has hitherto remained unexplored.
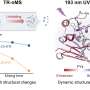
By combining protein X-ray crystal structures with supporting molecular biological and biochemical experiments, Dr. David Young and Dr. Brandán Pedre together with international collaborators and co-workers of the Messens lab have provided new insight into this question. They have uncovered the precise H2O2 binding site and the conformational changes that OxyR uses to bind to DNA and stimulate the regulation of the cellular H2O2 concentration.
"Previously, the H2O2-induced structural change of OxyR has led to the development of fluorescence-based genetically encoded H2O2 sensors, offering a way to visualize compartment-specific endogenous H2O2 in real time in living cells in various pathological conditions," explains Dr. David Young (VIB-VUB). Brandán Pedre (VIB-VUB) adds: "This new insight in the structural details of the OxyR protein not only clarifies how the cell arms itself against H2O2 changes but will also enable us to create more sensitive and specific OxyR-based fluorescent biosensors. Such sensors will help us to better understand how aberrant H2O2 signaling leads to disease and, in the long run, identify new drug targets."
More information: Brandán Pedre et al. Structural snapshots of OxyR reveal the peroxidatic mechanism of H2O2 sensing, Proceedings of the National Academy of Sciences (2018). DOI: 10.1073/pnas.1807954115
Journal information: Proceedings of the National Academy of Sciences
Provided by VIB (the Flanders Institute for Biotechnology)