Understanding water's role in antibiotic resistance emergence and dissemination in Africa

Greater access to antibiotic drugs, together with their misuse and overuse, has accelerated the emergence of antibiotic resistant bacteria worldwide.
A new study now suggests that surface water dynamics are a crucial contributor to this growing global health concern.
Funded by the National Science Foundation, Kathleen Alexander and Claire Sanderson of Virginia Tech's College of Natural Resources and Environment, together with researchers from two other universities, have used the Chobe River, the only permanent surface water resource in northern Botswana, to explore antibiotic resistance emergence and dissemination.
Alexander's unique field site in Botswana encompasses Chobe National Park, home to the largest population of African elephants in the world, as well as two townships. More importantly, the site lacks commercial agriculture or large-scale medical facilities, both believed to be main contributors to antibiotic resistance.
The team's findings, published in Frontiers in Microbiology, suggest that factors beyond industrial farming and medical facilities may be significant contributors to the global problem of antibiotic resistance dissemination.
The research team conducted surface water testing at transect sites along the Chobe River from the protected landscape within Chobe National Park through the urban town of Kasane to the mixed-use area of Kazungula that incorporates land used for subsistence farming, wildlife corridors, and human settlement.
The area has an appreciable wet (November-March) and dry (April-October) season and is significantly influenced by a flood-pulse pattern that brings rain waters from the Angolan highlands thousands of miles away.
"There are a lot of stakeholders involved in combating antibiotic resistance, but current action plans focus solely on the health care and agriculture sectors as the most important disseminators of antibiotic resistant bacteria into the environment," said Sanderson, research associate in Virginia Tech's Department of Fish and Wildlife Conservation and lead author of the study.
"In Botswana, we have a unique study site where we don't have any commercial agricultural enterprises or an extensive medical facility, but we've still found widespread antibiotic resistance in the surface water, spanning both the protected and urban landscapes. Additionally, the antibiotic resistance profiles we see in the surface water match those in the wildlife and human populations that share this important water resource," she continued.
Alexander, professor in the Department of Fish and Wildlife Conservation and an affiliate of Virginia Tech's Fralin Life Science Institute, echoed that water is the critical tie-in for all of the factors contributing to antibiotic resistance in the Chobe region. "Water is the resource that joins everyone and everything across the landscape. It brings all the connections and factors together: animal, human, and environmental."
The study found that land use and season were both statistically significant predictors of the presence of antibiotic resistant bacteria in surface water, with the wet season resulting in a higher mean antibiotic resistance than the dry season. Maps of the wet and dry season samples showed that bacterial isolates in water had the highest resistance to antibiotics in the town of Kasane, with significantly lower rates in Chobe National Park and the mixed land-use area of Kazungula.
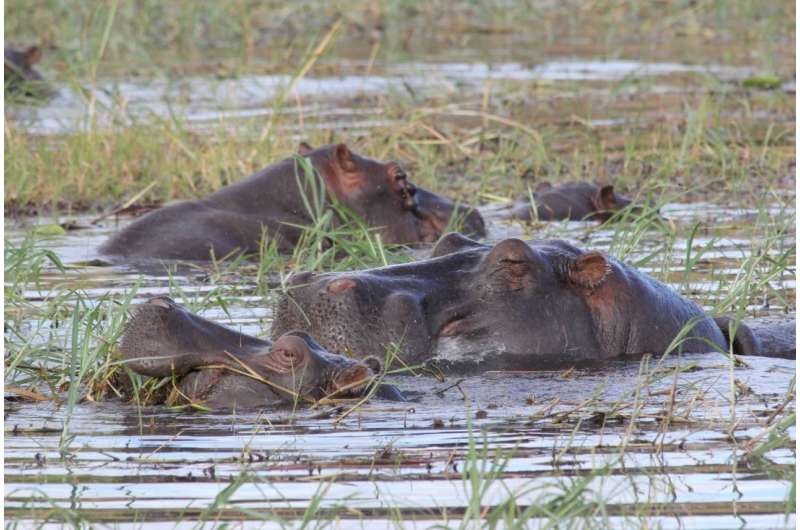
A previous study conducted by Alexander in her Botswana study site measured antibiotic resistance levels in fecal samples of water-associated animals (hippopotamus, crocodile, otter, and waterbuck) and non-water-associated animals (African elephant, banded mongoose, bushback, Cape buffalo, Chacma baboon, domestic cattle, giraffe, kudu, guineafowl, impala, leopard, sable, spotted hyena, vervet monkey, and warthog).
The study found that there was little to no antibiotic resistance in non-water-associated herbivores, but water-associated wildlife had higher levels of antibiotic resistance, indicating that the duration of time in the water and/or consumption of water-dwelling vegetation may play a critical role.
Alexander, who has been conducting research and collecting data in the area for more than 20 years, stressed that it is important to consider the issue of antibiotic resistance with a wider lens.
"Right now, antibiotic resistance is studied as a snapshot, but it changes spatially and temporally, and we need research that adjusts to those changes and advances our understanding of the environmental interactions between humans, animals, and the landscape," said Alexander, co-founder of the nonprofit Center for Conservation of African Resources: Animals, Communities, and Land Use (CARACAL) in Botswana.
The motivation for understanding antibiotic resistance is especially crucial in this area of Botswana, which claims some of the highest rates of HIV and tuberculosis in the region. "The need for cheap and effective antibiotics to help this vulnerable human population is extremely important," Sanderson added.
One avenue for future research is the idea of using native wildlife species in the Chobe region as surveillance systems to monitor changes in antibiotic resistance.
"We may be able to use the banded mongoose as a sentinel surveillance system because these animals live in close association with humans, and we have shown that antibiotic resistance is shared among water, humans, and wildlife in this region," Alexander said. "By routinely collecting feces from this species and testing the samples for antibiotic resistant bacteria, we will be able to monitor antibiotic resistance levels in the environment over time using noninvasive and time- and cost-effective techniques."
Sanderson and Alexander stress that understanding the causes and factors of antibiotic resistant bacteria emergence and dissemination demand a thorough comprehension of the interplay of environmental factors, human contributions, and patterns of human and wildlife interaction with this crucial water resource.
Provided by Virginia Tech