Unleashing TIGER on small RNAs
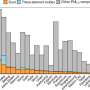
Efforts to explore the landscape of small RNAs (sRNAs)—short RNA molecules that are poorly understood—often use high-throughput sequencing (sRNA-seq). These efforts are hampered by a lack of tools to identify, quantify and analyze all the different sRNAs in sRNA-seq datasets.
Kasey Vickers, PhD, and colleagues have now developed a new analytical method called TIGER (Tools for Integrative Genome analysis of Extracellular sRNAs). To demonstrate the power of this tool, they performed sRNA-seq on mouse lipoproteins, bile, urine and livers. In the resulting datasets, TIGER was able to identify approximately 60 percent of sRNAs on lipoproteins and more than 85 percent of sRNAs in bile, urine and liver.
The researchers found that the majority of sRNAs on lipoproteins are non-host sRNAs derived from bacterial sources in the microbiome and environment.
The study, reported in the Journal of Extracellular Vesicles, revealed novel lipoprotein and biofluid sRNAs and showed that TIGER is well-suited to the study of sRNAs.
More information: Ryan M. Allen et al. Bioinformatic analysis of endogenous and exogenous small RNAs on lipoproteins, Journal of Extracellular Vesicles (2018). DOI: 10.1080/20013078.2018.1506198
Provided by Vanderbilt University