The mechanisms of genetic diversification in Candida albicans
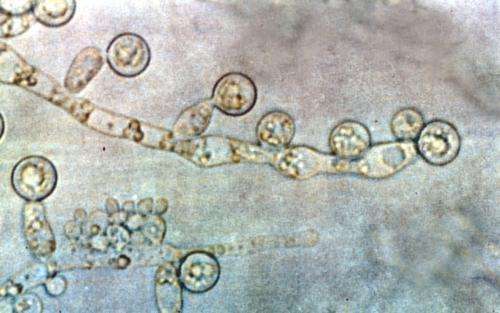
Candida albicans is one of the most formidable fungal species infecting humans. Investigating the structure and reproduction methods of pathogenic populations can reveal how they emerge and spread. A team of scientists has sequenced and analyzed the genomes of 182 strains of C. albicans from around the world. They confirmed the clonal reproduction of this C. albicans, and also showed that parasexual reproduction, previously only observed in a laboratory setting, contributes to its genetic diversity, and therefore also to its ability to adapt to new environments and rid itself of deleterious mutations.
There are 5 million fungal species, but only a few hundred can cause disease in humans. Candida albicans is one of the most formidable of these species. It belongs to one of the four genera of pathogenic fungi responsible for high mortality rates in humans and is the second most common agent of opportunistic fungal infection in the world. Candida albicans is part of the human gut microbiota (a commensal fungus) but it also causes mucosal infections in healthy individuals and severe opportunistic infections in those with weakened immune defenses (immunocompromised individuals and patients who have received organ transplants, undergone serious surgery or suffered major trauma).
Understanding how pathogens emerge and spread involves analyzing the structure of their populations. Several studies have shown the importance of population genetics in shedding light on the emergence of new diseases, such as white-nose syndrome in bats in North America, which is caused by a fungus and is devastating entire bat populations. In this case, population genetics studies revealed that a fungus of European origin, which is harmless in European bat populations, invaded North America via clonal expansion.
Scientists in the Biology and Fungal Pathogenicity Unit at the Institut Pasteur and INRA, in collaboration with 12 other teams, sequenced and analyzed the genomes of 182 strains of C. albicans isolates, either commensal or responsible for superficial or invasive infections, from around the world. This is the largest population genomics study carried out on the pathogen to date. It confirms the primarily clonal reproduction of this human pathogen. But it also shows traces of introgression in the genome of some strains, indicating the possibility of genetic exchanges between strains in nature and reflecting parasexual reproduction, independent from meiosis, which had previously only been observed in a laboratory setting, or sexual reproduction, hitherto unknown for C. albicans. C. albicans' use of (para)sexual reproduction is no doubt crucial for it to generate genetic diversity and adapt to new environments quickly, as well as to rid itself of the deleterious mutations that build up during clonal reproduction and which, if they are not eliminated, would lead to the species' extinction.
More information: Jeanne Ropars et al, Gene flow contributes to diversification of the major fungal pathogen Candida albicans, Nature Communications (2018). DOI: 10.1038/s41467-018-04787-4
Journal information: Nature Communications
Provided by Pasteur Institute