Your smartphone could help to speed up cancer research while you sleep
A new research project aims to speed up the delivery of personalised cancer treatments by using smartphones to crunch data while their owners sleep.
Researchers at Imperial College London are working with the Vodafone Foundation to recruit people to donate the power of their smartphones and run a simple app, which can help to carry out research overnight.
The project will harness the processing power of thousands of smartphones which, when combined, can analyse huge volumes of data in less time than it would take a supercomputer, and at a fraction of the cost of cloud computing platforms.
Smartphones contain a huge amounts of computing power needed to run everything from email to music and video streaming apps, but they are mostly dormant while users plug in their phones to charge overnight.
Drugs project
As part of the "DRUGS" (Drug Repositioning Using Grids of Smartphones) project, a team led by Dr. Kirill Veselkov in the Department of Surgery & Cancer at Imperial has designed an algorithm that breaks down enormous datasets into small chunks which can be analysed.
Users download the DreamLab app onto their phone and run it for six hours overnight as the phone charges.
While they sleep, the app downloads a small bite-sized packet of data – roughly 5 MB in size – and uses the phone's processors to run millions of calculations, before uploading the results and clearing the data.
The app has already been used by researchers in Australia to crunch data for pancreatic cancer, but this project will be the first time it has been used in the Europe.
By using the crowd-based approach to work on publicly available data on cancer genes and drug interactions, the Imperial researchers hope to significantly speed up cancer research by identifying new combinations of drugs that may be more effective in fighting cancers in individual patients.
Ultimately, using this mobile cloud-based computing approach could drastically reduce the time taken to analyse such vast amounts of data. A desktop computer with an eight-core processor running 24-hours a day would take 100 years to process the data. But a network of 100,000 smartphones running six hours per night could do the job in just three months.
Professor David Gann CBE, Vice President (Innovation) at Imperial College London, said: "Through harnessing distributed computing power, DreamLab is helping to make personalised medicine a reality.
"This project demonstrates how Imperial's innovative research partnerships with corporate partners and members of the public are working together to tackle some of the biggest problems we face today, generating real societal impact."
Andrew Dunnett, Director at the Vodafone Foundation, said: "We want to encourage people across the UK to become overnight heroes in the fight against cancer.
"Vodafone customers and users of other networks can use their smartphones to enable huge volumes of data to be processed.
"This means that Imperial College London will be able to speed up much-needed cancer research in the UK."
Genetic mutations and cancer
The hope is that rather than a trial and error approach of testing cancer drug combinations to see which work best for a patient, data-led approaches such as this could help to identify combinations of drugs to use based on the genotype of the cancer itself.
Dr. Veselkov said: "We are currently generating huge volumes of health data around the world every day, but just a fraction of this is being put to use. By harnessing the processing power of thousands of smartphones, we can tap into this invaluable resource and look for clues in the datasets.
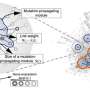
Using publicly available, anonymised research data drawn from several sources, the Imperial team designed an algorithm to explore how these mutations potentially interact with other genes."Ultimately, this could help us to make better use of existing drugs and find more effective combinations of drugs tailored to patients, and improving treatments."
The algorithm tests all possible combinations of gene-gene interactions based on existing data from almost one million cancer samples around the world, representing between 30,000 and 50,000 unique cancer mutations.
This type of analysis helps to 'stratify' patients into groups – ie those which have similar mutations, or gene-interactions in common – based on the genetic makeup of their cancer, rather than classification based on what type of cancer they have, or how advanced it is.
Exploring the connections
The algorithm also crunches data for known interactions for thousands of registered drugs, showing which genes and proteins they interact with – for instance, some drugs may downregulate a protein which is known to be upregulated by a specific cancer mutation.
This data includes cancer drugs as well as repurposed drugs initially designed for another indication. One example is sildenafil citrate, a compound which was originally designed as a cardiovascular drug but is now used to treat erectile dysfunction.
By overlaying the results, the team hope to build up a clearer picture of which drug combinations are most suited to patients, based on their mutations.
In future, the approach could include pharmacological data on the effects of the drugs and drug-drug interactions, to generate even more detailed information on which drug combinations and doses may work best for patients.
Dr. Veselkov added: "Every cancer patient in the UK could have DNA tested within the next decade. The challenge is how to use patients own data for more personalized therapy selection.
"In this exciting collaborative project with Vodafone, we have implemented and deployed the machine-learning algorithms on mobile phones to simulate the effect of cancer mutations and drugs on intracellular molecular circuits.
"The outcomes will shed light on possible multi-drug therapies against disrupted molecular networks rather than specific mutations in individual cancer patients."
More information: mediacentre.vodafone.co.uk/pressrelease/dreamlab/
Provided by Imperial College London