Physicists with green fingers estimate tree spanning rate in random networks
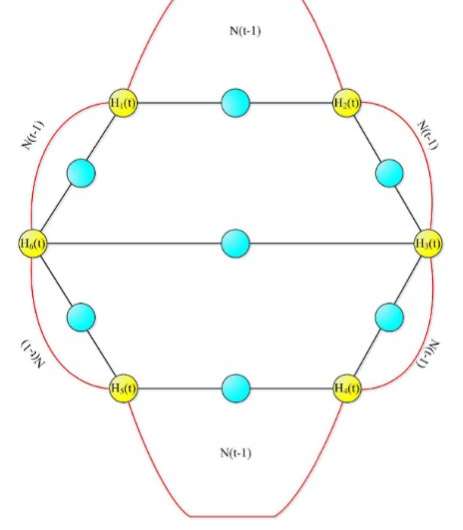
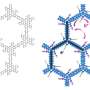
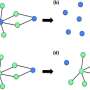
Networks are often described as trees with spanning branches. How the tree branches out depends on the logic behind the network's expansion, such as random expansion. However, some aspects of such randomly expanding networks are invariant; in other words, they display the same characteristics, regardless of the network's scale. As a result, the entire network has the same shape as one or more of its parts.
In a new study published in EPJ B, Fei Ma from Northwest Normal University in Lanzhou, Gansu Province, China, anc colleagues calculate the total number of spanning trees in randomly expanding networks. This method can be applied to modelling scale-free network models, which, as it turns out, are characterised by small-world properties. This means, for instance, that members of the network only exhibit six degrees of separation, like most people in our society.
Previously, a number of network models were based on graphs consisting of an aggregation of vertices with connecting edges. But they were not sufficient to model real-life networks, like networks of social media users. Instead, complex networks, where the network is created randomly, have become the mainstays of computer science and modern discrete mathematics. Using data from real-life networks, and drawing on the experience gained from artificial networks created to account for specific functions, the authors design more realistic models that are more complex than their predecessors.
In this study, the authors focus on developing a recursive method for calculating the number of spanning trees in a network, which is particularly helpful for predicting its capacity to tolerate faults that occur at random. Being able to find the number of spanning trees in network models has implications for various scientific fields, such as applied mathematics, theoretical computer science, physics and chemistry.
More information: Fei Ma et al, A recursive method for calculating the total number of spanning trees and its applications in self-similar small-world scale-free network models, The European Physical Journal B (2018). DOI: 10.1140/epjb/e2018-80560-8
Journal information: European Physical Journal B
Provided by Springer