April 12, 2018 report
Team creates detailed map of genetic evolution of Saccharomyces cerevisiae
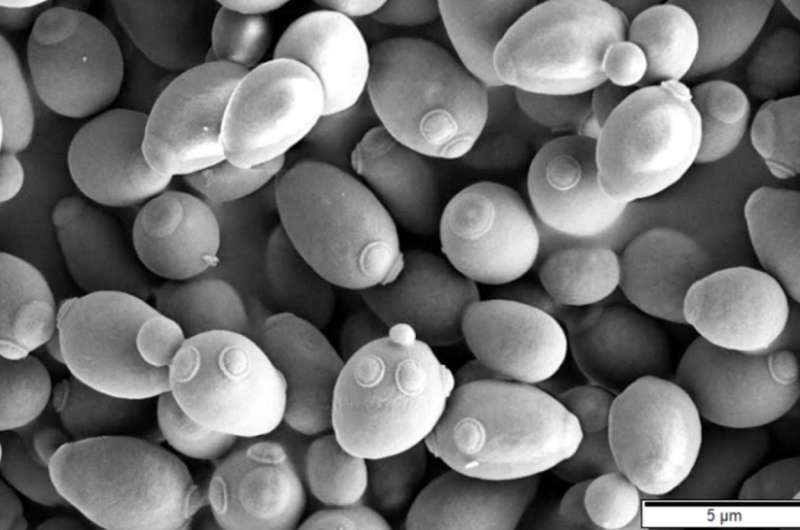

A team of researchers with members from several institutions in France has created a detailed map of the genetic evolution of Saccharomyces cerevisiae. In their paper published in the journal Nature, the group describes their analyses of the common yeast and what they found.
Saccharomyces cerevisiae, as the researchers note, has been used extensively by humans for a very long time—it is the yeast used to make bread, wine and beer. It is also widely used by research scientists because its genome has been sequenced and can be easily exploited. In this new effort, the researchers sought to learn more about yeast's history by studying its genetic makeup.
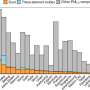
Prior research has suggested that humans first began to use yeast after it was found on the skin of grapes, and it has been theorized that its use first began in China. To find out more about its history, the researchers collected samples from around the globe—yeast in different parts of the world has changed over time partially due to natural events and partially due to human intervention. Taking a genomic global survey of yeast allowed the researchers to create a map of the evolution of the fungus. The work consisted of performing whole-genome sequencing on 1,011 samples of yeast which yielded 1,625,809 high-quality reference-based SNPs.
The researchers report that theories of an out-of-China migration of the yeast appears to be true. They also found that yeast has undergone extensive change due to human intervention related to fermentation for making beer, wine and sake. They noted also that the number of chromosome sets (ploidy) in the yeast cells had an impact on fitness across all of those species tested. They further noted that those with a diploid state appeared to have an advantage over other ploidy types. And they found that yeast has suffered from a loss of heterozygosity, in which there are two distinct alleles of a given gene present. They suggest this has likely led to the wide variation between the samples they observed.
The team concludes by suggesting that their work has laid a foundation for future genome research efforts in addition to offering a detailed history of the common yeast.
More information: Genome evolution across 1,011 Saccharomyces cerevisiae isolates, Nature (2018) doi:10.1038/s41586-018-0030-5
Abstract
Large-scale population genomic surveys are essential to explore the phenotypic diversity of natural populations. Here we report the whole-genome sequencing and phenotyping of 1,011 Saccharomyces cerevisiae isolates, which together provide an accurate evolutionary picture of the genomic variants that shape the species-wide phenotypic landscape of this yeast. Genomic analyses support a single 'out-of-China' origin for this species, followed by several independent domestication events. Although domesticated isolates exhibit high variation in ploidy, aneuploidy and genome content, genome evolution in wild isolates is mainly driven by the accumulation of single nucleotide polymorphisms. A common feature is the extensive loss of heterozygosity, which represents an essential source of inter-individual variation in this mainly asexual species. Most of the single nucleotide polymorphisms, including experimentally identified functional polymorphisms, are present at very low frequencies. The largest numbers of variants identified by genome-wide association are copy-number changes, which have a greater phenotypic effect than do single nucleotide polymorphisms. This resource will guide future population genomics and genotype–phenotype studies in this classic model system.
Journal information: Nature
© 2018 Phys.org