Democratizing single-cell analysis
Scientists at the Allen Institute and the University of Washington have developed a new low-cost technique for profiling gene expression in hundreds of thousands of cells. Split Pool Ligation-based Transcriptome sequencing (SPLiT-seq) is a scalable technique for characterizing RNA in individual cells that can be used to identify the various cell types found in the brain and other tissues. The research is published this week in the journal Science.
"To understand any multicellular biological system, we need to understand its building blocks, its cells," said Bosiljka Tasic, a scientist at the Allen Institute. "The brain contains many cells of different shapes and sizes, but a universal approach to characterize and classify them does not exist. Characterizing RNA in single cells is a way to classify cells, but the existing methods are expensive and require specialized equipment. Since SPLiT-seq can be done with basic equipment in any lab it may lead to wider adoption and standardization of single cell transcriptomics."
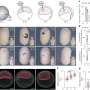
Previously developed sequencing techniques require that each cell is isolated before its RNA is labeled with a genetic barcode. With SPLiT-seq, cells are never isolated. Instead, the cell is used as an isolation compartment for its own RNA. Pools of cells are split into multiple groups where a barcode is added to their RNA. This process is repeated multiple times to give the RNA of each cell a unique combinatorial barcode. This allows researchers to profile the gene expression of many cells at the same time and classify them into types.
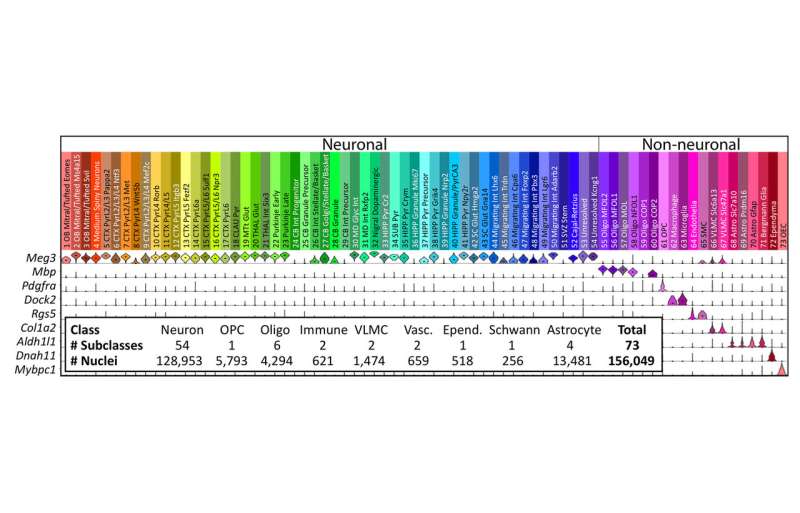
"For this project, we used SPLiT-seq to profile more than 150,000 cells in the developing mouse brain and spinal cord, and we defined many new molecular markers for specific cell type," said Georg Seelig, a synthetic biologist at the Department of Electrical Engineering and the Paul G. Allen School of Computer Science and Engineering at the University of Washington. "Our experiments cost just a penny per cell. SPLiT-seq significantly lowers the barrier for labs that want to do single cell profiling but can't afford expensive, specialized equipment."
"Many diseases, such as Alzheimer's and Parkinson's, begin with dysfunction in certain cell types," said Tasic. "Insights into what those types are and which genes have gone awry will lead to a better understanding of how these diseases start and progress, and how to treat them."
Journal information: Science
Provided by Allen Institute for Brain Science