January 5, 2018 report
DropSynth, a one-pot approach to gene synthesis

A team of researchers at the University of California has found a way to synthesize multiple genes from a group of microarray-generated oligonucleotides. In their paper published in the journal Science, the group describes their technique, called DropSynth, how well it works, and its drawbacks.
Synthesizing genes has become so popular that there are now companies that do it for a living, but it is still an expensive proposition—current methods require sewing small strands into sequences one at a time after they have been created. In this new effort, the researchers report a one-pot approach to gene synthesis that could lead to lowering costs.
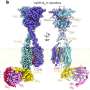
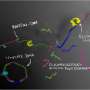
Currently, gene synthesis is done using a microarray, producing DNA oligonucleotides, which then have to be sewn together—the team at UoC also started with a microarray, but they first gave the oligonucleotides an identifying length of gene sequence, which they describe as a "barcode." Next, they added microbeads with complimentary barcodes that could pull matching oligonucleotides from a pool of different kinds of oligonucleotides. The result was a pool of microbeads, each of which had a small group of attached same-type oligonucleotides. The team then enclosed each of the microbeads (and its group of oligonucleotides) in an emulsion droplet using what they describe as an oligonucleotide and oil mixture vortex—for 30 seconds. After that, enzymes induced all of the oligonucleotides in a single droplet to merge (via polymerase cycling assembly), resulting in a desired sequence. The sequences were then removed from the emulsion ready for use.
The process is better than conventional methods, the team suggests, because it offers access to a gene pool at essentially the same cost as an oligonucleotide pool—it does not add much. Unfortunately, there is a pretty big downside. The process, they note, is "messy," because it is only approximately 5 percent efficient—not nearly good enough for use in manufacturing processes. On a more positive note, the method could be used to create large libraries of genes for various purposes, at very little cost. It could also be used in research efforts, particularly those involved in protein design.
More information: Calin Plesa et al. Multiplexed gene synthesis in emulsions for exploring protein functional landscapes, Science (2018). DOI: 10.1126/science.aao5167
Abstract
Improving our ability to construct and functionally characterize DNA sequences would broadly accelerate progress in biology. Here, we introduce DropSynth, a scalable, low-cost method to build thousands of defined gene-length constructs in a pooled (multiplexed) manner. DropSynth uses a library of barcoded beads that pull down the oligonucleotides necessary for a gene's assembly, which are then processed and assembled in water-in-oil emulsions. We use DropSynth to successfully build >7000 synthetic genes that encode phylogenetically-diverse homologs of two essential genes in E. coli. We tested the ability of phosphopantetheine adenylyltransferase homologs to complement a knockout E. coli strain in multiplex, revealing core functional motifs and reasons underlying homolog incompatibility. DropSynth coupled with multiplexed functional assays allow us to rationally explore sequence-function relationships at unprecedented scale.
Journal information: Science
© 2018 Phys.org