The future of crop engineering
Photosynthesis is the process underlying all plant growth. Scientists aim to boost photosynthesis to meet the increasing global demand for food by engineering its key enzyme Rubisco. Now, researchers at the Max Planck Institute of Biochemistry have succeeded in producing functional plant Rubisco in a bacterium. This allows genetic engineering of the enzyme. The study could one day lead to better crop yields and plant varieties with increased water-use efficiencies or enhanced temperature resistance. The results were published in Science.
The world's population is predicted to exceed 9 billion in 2050. With more mouths to feed, there is a pressing need for improved food output. To meet the global demand for food, scientists aim to increase the efficiency of photosynthesis and therefore crop productivity.
Boosting photosynthesis
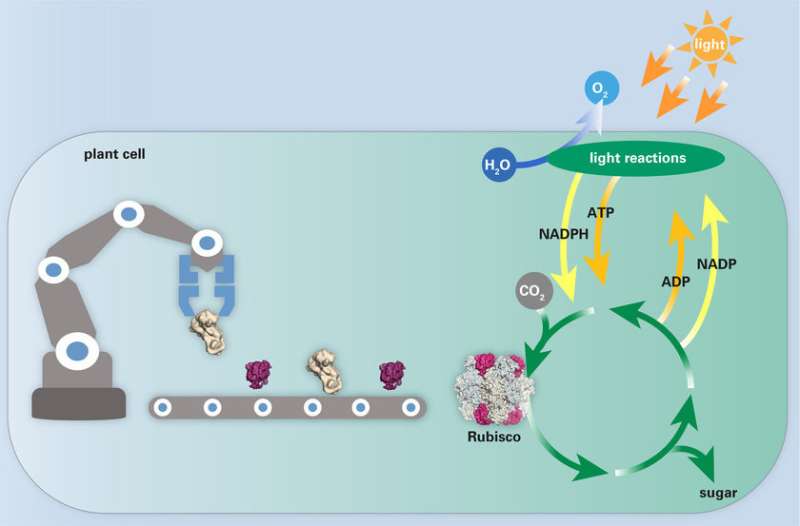
Photosynthesis is the fundamental biological process that underlies all plant growth and supports life on Earth. Plants use the energy of sunlight to convert carbon dioxide (CO2) and water to sugar and oxygen (O2). The critical enzyme in this process is Rubisco. Rubisco catalyses the first step in carbohydrate production in plants, the fixation of CO2 from the atmosphere. In doing so, plants utilize CO2 to build biomass and produce the required energy for growth. However, Rubisco is an inefficient enzyme as it captures CO2 slowly. Competing reactions with O2 further impair Rubisco's catalytic efficiency. For these reasons, Rubisco often limits the rate of photosynthesis and ultimately plant growth, making Rubisco a hot target for genetic engineering.
Engineering of plant Rubisco, and photosynthesis, would be enhanced by functional expression of the enzyme in alternative hosts. So far, however, scientists failed to produce an enzymatically active form of plant Rubisco in a bacterial host – a goal that has been sought after for many decades. A team led by Manajit Hayer-Hartl, head of the research group "Chaperonin-assisted Protein Folding", has now identified the requirements for expressing and assembling plant Rubisco in a bacterium. Their findings are expected to greatly accelerate efforts to improve photosynthesis through Rubisco engineering.
The Rubisco assembly line
The Rubisco enzyme consists of eight large and eight small subunits. Protein folding of the large subunits is assisted by specific chaperonins, macromolecular folding cages, in which the newly synthesized proteins can assume their proper functional conformation. After folding, multiple additional helper proteins (chaperones) assist in the proper assembly of the subunits into the large enzyme complex.
The researchers generated functional plant Rubisco in a bacterial host by simultaneously expressing plant chaperones and Rubisco in the same cells. This not only enables the scientists to understand the complex assembly pathway of Rubisco, but to modify the Rubisco gene in order to improve Rubisco's properties. Once they have obtained a Rubisco variant with a desired trait, they can insert the modified gene back into the plant cells. This is a key-step towards improving photosynthesis through Rubisco engineering. "The bacterial expression system resembles an assembly line for cars. Whereas previously every optimized variant of Rubisco had to be painstakingly expressed in a transgenic plant, which takes a year or more to generate - like building a car by hand - we can now make hundreds or thousands of Rubisco variants in days or weeks. It is like building cars in an automated assembly line", explains Hayer-Hartl.
Superior Rubisco variants
Genetic engineering facilitates efforts to generate Rubisco variants with improved functional properties. This might not only lead to the much-needed increase in crop yields, but also plant varieties with increased water-use efficiencies or enhanced temperature resistance - properties that are of special importance in the light of global warming and increasing water scarcity.
More information: H. Aigner et al. Plant RuBisCo assembly inE. coliwith five chloroplast chaperones including BSD2, Science (2017). DOI: 10.1126/science.aap9221
Journal information: Science
Provided by Max Planck Society





















