Biophysics study makes exciting advancements for the future of DNA sequencing
A Northeastern research team has developed new technology that optimizes DNA sequencing using nanophysics and electric currents. In a paper published in Nature Nanotechnology, Northeastern Professor of Biological Physics Meni Wanunu, in partnership with Pacific Biosciences, a biotechnology company with a focus on DNA sequencing, developed a method for loading DNA into sequencing wells with orders of magnitude higher efficiencies.
"Apart from being a multi-billion dollar a year market, DNA sequencing is one avenue where incremental improvements in research, like discovery of a new gene, for example, can have immediate clinical consequences," said Wanunu.
Our human DNA is a genome composed of 23 pairs of chromosomes, which breaks down into six billion pieces that all come together to give each person their unique characteristics and properties. While we have the ability to sequence important parts of the genome, the ability to know the entire sequence has the potential to make huge strides in the area of understanding and predicting disease, and more importantly, to personalize medicine.
"Right now, piecing together the entire sequence through traditional methods is like stitching together a giant puzzle, and the error rate can get so enormous that after the first few hundred bases, the sequence is gibberish," said Wanunu. "That's why there's a fundamental limit for second-generation sequencing methods, which we want to move past."
This is why technology has evolved to bring forward a new method for sequencing DNA: single-molecule sequencing.
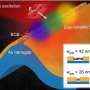
Pacific Biosciences has developed an optical technology for single-molecule DNA sequencing that relies on nano-wells. These wells localize the sequencing signal and allow single molecule sequencing to be carried out. However, the methods used by the company to load the DNA into the wells favors shorter DNA molecules, rather than longer ones.
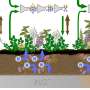
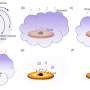
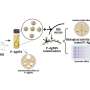
Wanunu's lab has redesigned the wells to incorporate nanopores at their bases, which allows them to attract larger segments of DNA using an electric field. By simply applying a voltage, the charged DNA molecules efficiently enter the wells, and longer DNA molecules become preferred over shorter ones.
"Large DNA molecules need just a small push to get into the sequencing volume, but once we apply this force, we can capture enormous sample fragments easily. The system will enable totally new sequencing experiments," said Joe Larkin, first author of this paper.
To further this research, Wanunu and his lab are working on preparing this technology for more large-scale use, specifically with equipment at Pacific Biosciences. The team is testing a porous substrate to replace the metal wells currently being used to attract and sequence DNA. As they continue in this research, Wanunu hopes to even further increase the fundamental length of DNA that can be sequenced.
"We would like to have a platform, some day, that sequences every nucleic acid molecule in a single cell, without the need for making many copies of these molecules prior to sequencing, just reading the native DNA," Wanunu said.
More information: Joseph Larkin et al, Length-independent DNA packing into nanopore zero-mode waveguides for low-input DNA sequencing, Nature Nanotechnology (2017). DOI: 10.1038/nnano.2017.176
Journal information: Nature Nanotechnology
Provided by Northeastern University