Colon cancer nuclear pore dynamics are captured by HS-AFM
One of the key reasons for cancer mortality is the highly invasive behaviour of cancer cells, which is often due to aggressive metastasis. Metastasis is facilitated by various growth factors and cytokines secreted from cells of the immune system, which operate through various signaling pathways. Remarkably, these signaling pathways enter the nucleus through the nuclear pore complex (NPC), which is supposed to act as a doorkeeper to the nucleus. NPC is, in fact, a nanomachine consisting of multiple copies of about 30 different proteins, collectively called nucleoporin.
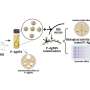
Although small molecules are able to go through the nuclear pores rather freely, molecules larger than 40 kDa could do so effectively only by binding to specific transporter proteins that interact with FG-Nups (nucleoporins have repeating units of two amino acids phenylalanine (F) and glycine (G), which are the tentacle proteins having specific and selecting roles in pore transportation. Although different models are proposed, how FG-Nups participates in the nucleus-cytoplasm transport remains largely unknown. Nonetheless, the concomitant assessment of nanoscopic structures and dynamics has been technically unfeasible, a situation prevailing throughout cell biology research. The direct visualization of NPC dynamics at nanoscale resolution was thought to be impossible.
The research team of Kanazawa University investigated this important issue and obtained the groundbreaking results by combined high-resolution live cell imaging, electron microscopy, and high-speed AFM (HS-AFM) which is developed by themselves to investigate the native nanoscopic spatial and temporal dynamics in NPC structures in the colon cancer cells.
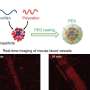
First, they generated NPC stable cell lines expressing GFP (green fluorescent protein) and confirmed by fluorescent microscopy. Next, they isolated the highly purified nuclear envelope which was confirmed by the use of negative stain electron microscopy and confocal microscopy. Then, they started the observation of spatiotemporal changes at millisecond and nanometer scale of native state NPC structure in colon cancer cells by combining high resolution live cell imaging and electron microscopy. Notably, they performed the observation of living nuclear envelope and nuclear pores using HS-AFM.
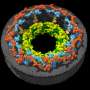
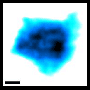
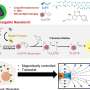
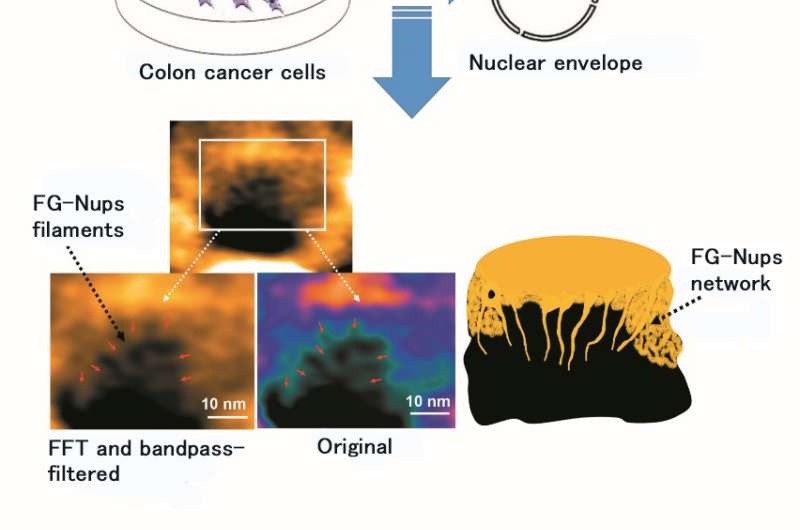
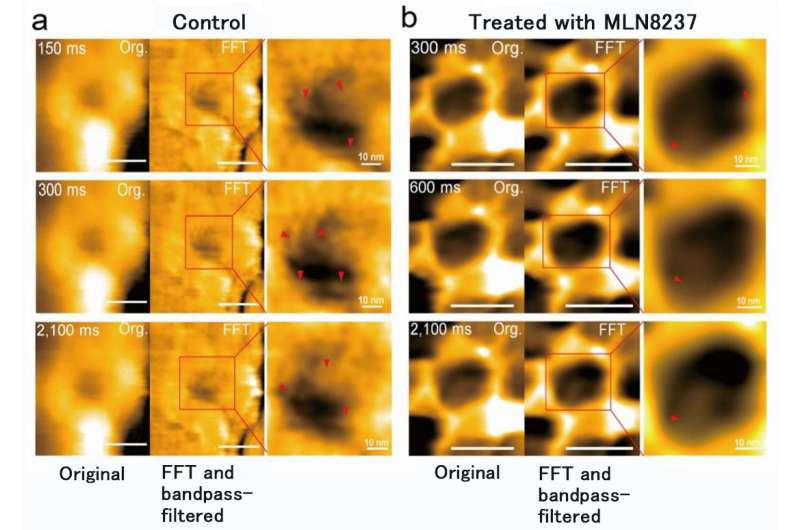
The research team of Kanazawa University was, indeed, successful in imaging the dynamics of NPC proteins in cancer cells, which are the building blocks of the nuclear pore (Figure 1). MLN8237/alisertib, an apoptotic and autophagic inducer, is currently undergoing several cancer clinical trials. This drug was reported to inhibit nucleoporin expression and activities. The researchers visualized native and drug-treated FG-Nups by HS-AFM. In particular, the extended and retracted FG-Nups having a spider cobweb appearance were lost in drug-treated samples (Figure 2). The research team concluded that via HS-AFM, they visualized the deformation and loss of FG-Nups nuclear pore barrier, which might be the first nano dying code discovered in the world.
The present study by the research team of Kanazawa University enabled visualization of structure and dynamics of the nuclear membrane pore at nanometer scale, and it is shown that deformation and loss of the nuclear membrane pore barrier would be one of the dying codes of cancer cells. These findings stand for a new paradigm in our understanding of nuclear transport, which has, up to this point, remained an enigmatic problem throughout the nano-medicine and cell biology field. Current findings are based on the crowning bio-imaging technology developed at Kanazawa University. This study has medical applications including acting as a novel "nano-endoscopy" to visualize intra-cellular organelles such as the nucleus and nuclear pores, and molecular dynamics in cancer cells and other diseases.
More information: Mahmoud Shaaban Mohamed et al, High-Speed Atomic Force Microscopy Reveals Loss of Nuclear Pore Resilience as a Dying Code in Colorectal Cancer Cells, ACS Nano (2017). DOI: 10.1021/acsnano.7b00906
Journal information: ACS Nano
Provided by Kanazawa University