Unveiling how nucleosome repositioning occurs to shed light on genetic diseases
A research group led by Hitoshi Kurumizaka, a professor of structural biology at Waseda University, unveiled the crystal structure of an overlapping dinucleosome, a newly discovered chromatin structural unit. This may explain how nucleosome repositioning occurs and provide valuable information for developing drugs to treat genetic diseases.
This research is published in Science.
Humans develop from a single cell, which divides repeatedly to form organisms. These divided cells all carry the same genetic information and differentiate to form tissues with different functions. When a hand is formed, for example, only the genes needed to form a hand are "switched on," and all the other genes other than those used to form a hand remain switched off. Epigenetics is the field of research that aims to explain this so-called genetic switch, which controls the expression of genes that compose the structure of each organism.
A DNA string stores all of a person's genetic information and measures 2 meters long. This long of DNA must be packed in a nucleus of only about 10 micrometers in diameter. To do so, the DNA is folded neatly into a structure called chromatin. To decipher the genetic information in this condition, the chromatin structure is dynamically modified for reading. This change in chromatin structure regulates the differences in genes being read and differentiate phenotypic variations of cells.
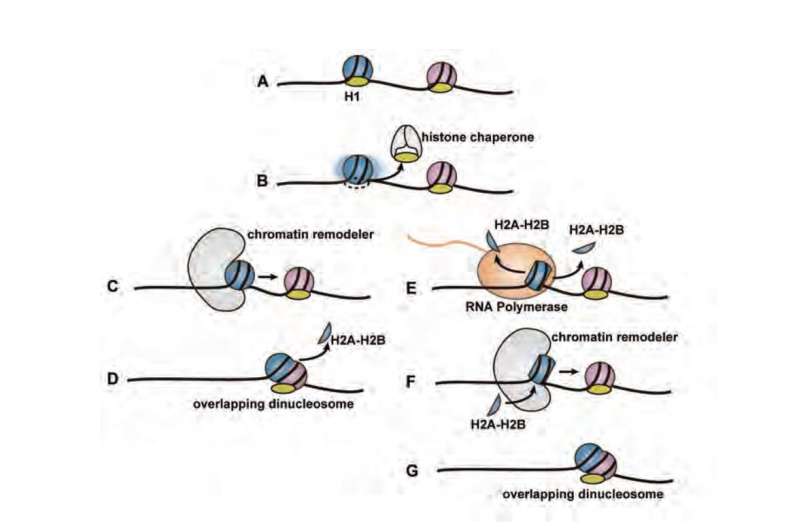
A chromatin is composed of four histone proteins with DNA wrapped around a chain of spools known as nucleosomes. When reading genetic information, a chromatin structure is modified, so that the DNA packed into the chromatin becomes easier to read by repositioning the nucleosome near the point where the reading starts. Then, an RNA polymerase, an enzyme that transcribes the genes, starts reading the DNA from where the nucleosome was moved. This phenomenon of repositioning is called nucleosome remodeling. It was hypothesized that in nucleosome remodeling, nucleosomes collide and form a chromatin structural unit known as an overlapping dinucleosome (Figure 01). The formation of an overlapping dinucleosome through nucleosome remodeling was thought to be crucial for regulating the genetic switch, but its existence and actual structure were elusive.
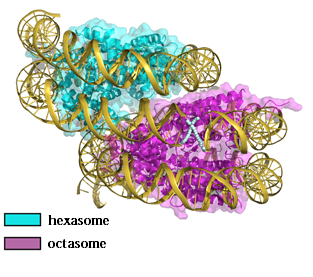
Professor Kurumizaka's research group determined the three-dimensional structure of an overlapping dinucleosome at atomic resolution.
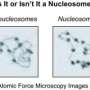
"Our team developed a method to reconstitute overlapping dinucleosomes in vitro and to prepare them in large quantities with high purity. We succeeded in crystallizing the purified overlapping dinucleosomes (Figure 02), and using these, we conducted x-ray diffraction experiments at SPring-8, a large-scale synchrotron radiation facility."
These discoveries are promising for expanding research on the link between the overlapping dinucleosome and the genetic switch.
Moreover, mutations in nucleosome remodeling proteins have been found in various cancers, including ovarian and bladder cancers. This suggests that an incomplete formation of overlapping dinucleosome may trigger abnormalities in the genetic switch, turning normal cells into tumor cells. Accordingly, understanding the atomic structure of the overlapping dinucleosome may provide valuable information for understanding how the abnormal dynamics of chromatin are related to cancer, offering essential information for development in cancer drugs.
Professor Kurumizaka is intrigued to study the correlation between the overlapping dinucleosome formation and the genomic DNA functions in cells in the future.
More information: "Crystal structure of the overlapping dinucleosome composed of hexasome and octasome" Science (2017). science.sciencemag.org/cgi/doi … 1126/science.aak9867
Journal information: Science
Provided by Waseda University