Super resolution imaging helps determine a stem cell's future
Scientists at Rutgers and other universities have created a new way to identify the state and fate of stem cells earlier than previously possible.
Understanding a stem cell's fate—the type of cell it will eventually become—and how far along it is in the process of development can help scientists better manipulate cells for stem cell therapy.
The beauty of the method is its simplicity and versatility, said Prabhas V. Moghe, distinguished professor of biomedical engineering and chemical and biochemical engineering at Rutgers and senior author of a study published recently in the journal Scientific Reports. "It will usher in the next wave of studies and findings," he added.
Existing approaches to assess the states of stem cells look at the overall population of cells but aren't specific enough to identify individual cells' fates. But when implanting stem cells (during a bone marrow transplant following cancer treatment, for example), knowing that each cell will become the desired cell type is essential. Furthermore, many protein markers used to distinguish cell types don't show up until after the cell has transitioned, which can be too late for some applications.
To identify earlier signals of a stem cell's fate, an interdisciplinary team from multiple universities collaborated to use super-resolution microscopy to analyze epigenetic modifications. Epigenetic modifications change how DNA is wrapped up within the nucleus, allowing different genes to be expressed. Some modifications signal that a stem cell is transitioning into a particular type of cell, such as a blood, bone or fat cell. Using the new method, the team of scientists was able to determine a cell's fate days before other techniques.
"Having the ability to visualize a stem cell's future will take some of the questions out of using stem cells to help regenerate tissue and treat diseases," says Rosemarie Hunziker, program director for Tissue Engineering and Regenerative Medicine at the National Institute of Biomedical Imaging and Bioengineering. "It's a relatively simple way to get a jump on determining the right cells to use."
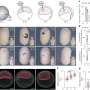
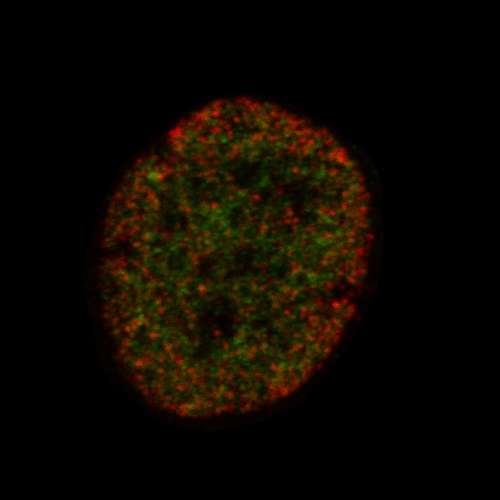
The approach, called EDICTS (Epi-mark Descriptor Imaging of Cell Transitional States), involves labeling epigenetic modifications and then imaging the cells with super resolution to see the precise location of the marks.
"We're able to demarcate and catch changes in these cells that are actually not distinguished by established techniques such as mass spectrometry," Moghe said. He described the method as "fingerprinting the guts of the cell," and the results are quantifiable descriptors of each cell's organization (for example, how particular modifications are distributed throughout the nuclei).
The team demonstrated the method's capabilities by measuring two types of epigenetic modifications in the nuclei of human stem cells cultured in a dish. They added chemicals that coaxed some of the cells to become fat cells and others to become bone, while another set served as control. Within three days, the localization of the modifications varied in cells destined for different fates, two to four days before traditional methods could identify such differences between the cells. The technique had the specificity to look at regional changes within individual cells, while existing techniques can only measure total levels of modifications among the entire population of cells.
"The levels are not significantly different, but how they're organized is different and that seems to correlate with the fact that these cells are actually exhibiting different fates," Moghe said. "It allows us to take out a single cell from a population of dissimilar cells," which can help researchers select particular cells for different stem cell applications.
The method is as easy as labeling, staining and imaging cells - techniques already familiar to many researchers, he said. As the microscopes capable of super resolution imaging become more widely available, scientists can use it to sort and screen different types of cells, understand how a particular drug may disrupt epigenetic signaling, or ensure that stem cells to be implanted won't transform into the wrong cell type.
More information: Joseph J. Kim et al, Optical High Content Nanoscopy of Epigenetic Marks Decodes Phenotypic Divergence in Stem Cells, Scientific Reports (2017). DOI: 10.1038/srep39406
Journal information: Scientific Reports
Provided by Rutgers University