Looking for life in all the right places—with the right tool
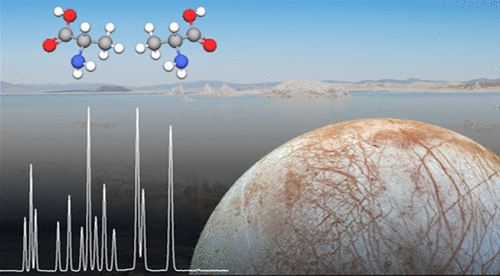
Researchers have invented a range of instruments from giant telescopes to rovers to search for life in outer space, but so far, these efforts have yielded no definitive evidence that it exists beyond Earth. Now scientists have developed a new tool that can look for signs of life with 10,000 times more sensitivity than instruments carried on previous spaceflight missions. Their report appears in the ACS journal Analytical Chemistry.
One path to finding life on other planets—or moons—involves looking for signature patterns of amino acids, which are organic molecules that are critical to life on Earth. But looking for these molecules on Mars or other planetary surfaces has been a major challenge. The Curiosity rover exploring Mars attempted to accomplish this, but the rover's experiments to identify organic chemicals in Martian samples were complicated by reactions with other materials in the samples. So Peter A. Willis, Jessica Creamer and Maria F. Mora set out to address this limitation.
The researchers created methods based on capillary electrophoresis to process soil or ice samples and detect 17 different amino acids simultaneously. This particular set of amino acids can be found in large quantities in biological and non-living samples, but in certain patterns, could serve as an indicator of life. The researchers validated their approach by analyzing samples from California's Mono Lake, an extremely salty body of water acting as a stand-in for briny water on Mars and on some moons. The methods detected the amino acids with 10,000 times the sensitivity of past approaches and identified three different biosignatures that were present. Willis, a member of the Europa Lander Science Definition Team, says that this type of technology is under consideration for future missions to ocean worlds like Europa and also Enceladus. The researchers say these are the best techniques yet to find signs of life on other worlds.
More information: Jessica S. Creamer et al. Enhanced Resolution of Chiral Amino Acids with Capillary Electrophoresis for Biosignature Detection in Extraterrestrial Samples, Analytical Chemistry (2016). DOI: 10.1021/acs.analchem.6b04338
Abstract
Amino acids are fundamental building blocks of terrestrial life as well as ubiquitous byproducts of abiotic reactions. In order to distinguish between amino acids formed by abiotic versus biotic processes it is possible to use chemical distributions to identify patterns unique to life. This article describes two capillary electrophoresis methods capable of resolving 17 amino acids found in high abundance in both biotic and abiotic samples (seven enantiomer pairs d/l-Ala, -Asp, -Glu, -His, -Leu, -Ser, -Val and the three achiral amino acids Gly, β-Ala, and GABA). To resolve the 13 neutral amino acids one method utilizes a background electrolyte containing γ-cyclodextrin and sodium taurocholate micelles. The acidic amino acid enantiomers were resolved with γ-cyclodextrin alone. These methods allow detection limits down to 5 nM for the neutral amino acids and 500 nM for acidic amino acids and were used to analyze samples collected from Mono Lake with minimal sample preparation.
Journal information: Analytical Chemistry
Provided by American Chemical Society