December 8, 2016 feature
Particles self-assemble into Archimedean tilings
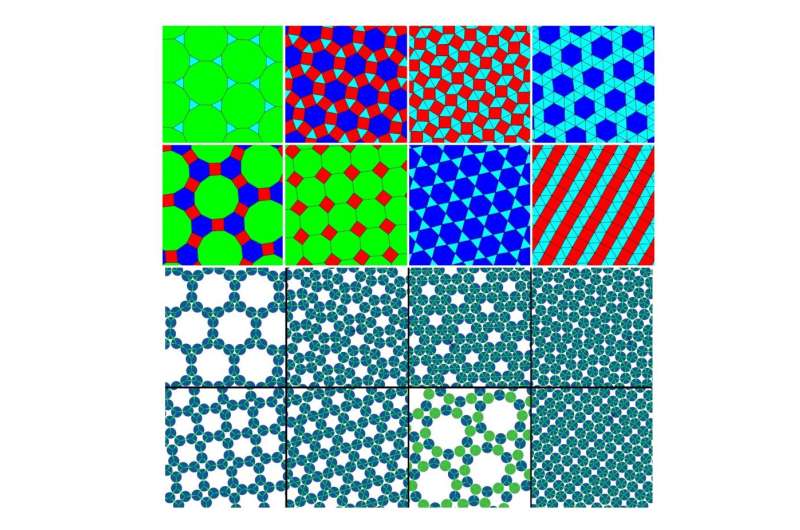

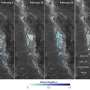
(Phys.org)—For the first time, researchers have simulated particles that can spontaneously self-assemble into networks that form geometrical arrangements called Archimedean tilings. The key to realizing these structures is a strategy called minimal positive design, in which both the geometry and the chemical selectivity of the particles is taken into account. The process has applications in molecular self-assembly, which could one day be used to build a variety of nanoscale technologies.
Stephen Whitelam, a researcher at the Molecular Foundry at Lawrence Berkeley National Laboratory, has published a paper on the minimal positive design strategy for self-assembling Archimedean tilings in a recent issue of Physical Review Letters.
Previously, researchers have successfully self-assembled particles into Platonic tilings, which are simpler arrangements consisting of regular periodic arrays of a single shape, such as squares, triangles, or hexagons. To do this, researchers use a strategy called positive design, in which the desired structure is promoted based on the particle geometry. When the particles are combined and cooled, they spontaneously self-assemble into Platonic tilings due to a variety of underlying chemical, physical, and thermodynamic interactions.
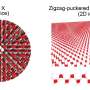
Self-assembling particles into the next-simplest arrangement, Archimedean tilings, is much more difficult. Archimedean tilings are composed of two or three different shapes, and only one type of vertex (so if you zoomed in on the intersection points, they would all look the same, having the same angles in the same order). There are eight types of Archimedean tilings, and the new design strategy can construct all eight of them.
The novel aspect of the new design strategy is the "minimal" element, which refers to chemical selectivity. Whitelam found that if you identify all of the interparticle interactions involved in a desired arrangement, and then select particles with only those interactions and no others, then under a simple cooling protocol the particles will self-assemble into the desired structure. The "positive" aspect of the strategy is that it works by promoting the desired structure, and does not require suppressing all of the many possible undesired structures.
The simulations also showed that, if chemical selectivity is not accounted for, then particles do not self-assemble into the Archimedean tilings, showing that the chemical selectivity is critical to realizing these structures.
"The results show that you need 'chemical specificity' of interactions to self-assemble certain simple, regular structures," Whitelam told Phys.org. "I wanted to write a paper about the amount of 'information' that you need to 'program' into a particle in order to allow it to self-assemble, in the presence of many copies of itself, into a desired structure.
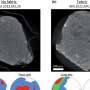
"The simplest way of tiling a plane is to cover it with triangles, or with squares, or with hexagons. These patterns are called the Platonic or regular Archimedean tilings. Other authors have shown that particles with certain geometric properties—with sticky patches at certain angles—can spontaneously form the networks that are equivalent to these tilings, meaning that if you draw lines between particle centers, then the picture you get looks like a tiling.
"The next simplest way to cover a surface is with combinations of two or three regular polygons, and these patterns are called the semi-regular Archimedean tilings (often just Archimedean tilings). Other researchers have used simulations to show that particles with the correct geometry alone probably can't self-assemble into such structures. My work confirms this fact, but shows that what does work is if the particle interactions are chemically specific, meaning that the sticky patches only stick to certain other sticky patches. In this way, particles avoid making lots of binding mistakes, and manage to find their way to the correct structure."
Interestingly, chemical selectivity is also used to control the interparticle interactions among biological particles, such as proteins and DNA.
"One aspect of this result is already widely known: researchers who use DNA nanotechnology routinely use DNA-mediated chemically specific interactions to make structures as complex or more complex than the Archimedean tilings," Whitelam said. "What surprised me is 1) that you need this strategy even for the Archimedean tilings (among the most simple regular structures), and 2) that all you need is this strategy (that is, chemical specificity is both 'necessary and sufficient'). I wanted to emphasize this fact because it did not seem obvious to me that one should need chemical specificity to make the networks equivalent to the Archimedean tilings."
Whitelam expects that the minimal positive design strategy should also work for self-assembling particles into other types of arrangements besides Archimedean tilings. The ability to self-assemble particles into various patterns could provide a tool for fabricating future nanoscale devices.
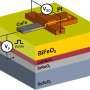
"Researchers would like to be able to control the patterns of molecules on surfaces for several reasons," Whitelam said. "One is that if you can control what sits on a surface and how it is organized then you can influence the chemical and physical properties of the surface. Another is that networks in two dimensions can be used to 'host,' or position, other molecules: imagine putting particles (such as metal nanoparticles, potentially useful in next-generation electronics devices) in the pores of the networks. It is important to be able to control the relative positions of such particles; being able to assemble any of the Archimedean tilings, or indeed any network you desired, would be a good way of doing this."
In the future, Whitelam plans to work with other scientists who come to The Molecular Foundry at Berkeley Lab, which is a US Department of Energy User Facility for nanoscience.
"My aim is to build on this work in order to collaborate with Foundry Users—scientists who come to the Foundry to do collaborative nanoscience—who make DNA-linked particles," he said. "We would like to work out how to design such particles in order to make new nanostructures 'to order.'
More information: Stephen Whitelam. "Minimal Positive Design for Self-Assembly of the Archimedean Tilings." Physical Review Letters. DOI: 10.1103/PhysRevLett.117.228003
Journal information: Physical Review Letters
© 2016 Phys.org