November 22, 2016 report
Sequencing of environmental DNA offers information on whale shark populations

(Phys.org)—A team of researchers with members from Denmark, the U.K. and Qatar was able to calculate whale shark populations in the Persian Gulf using only environmental DNA (eDNA) found in seawater samples. In their paper published in the journal Nature Ecology & Evolution, the team describes their analyses of sea water samples, what it revealed and the ways in which such information can be useful.
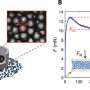
Whale sharks are the world's largest fish and survive by filtering material found in seawater (such as tuna eggs) and swallowing it. To learn more about these gentle giants, the researchers collected several small samples of ocean water from areas where whale sharks are known to reside. Then, they filtered the water looking for material likely to offer whale eDNA which would include feces, urine and bits of skin. By sequencing the whale material, the researchers were able to learn a great deal about the local whale population.
They were able to calculate the DNA mutation rate for the whale sharks—for example, by comparing what they found in the eDNA with samples obtained from other whale sharks at other times. From that, they were able to calculate the likely population size in the local area. Using the local population number, they were able to calculate the likely population size for the entire Indo-Pacific. The analysis also revealed that the whale sharks in the Indo-Pacific are genetically distinct from those living in the Atlantic Ocean.
Collecting and analyzing eDNA has become a useful tool for studying animals in a new way—its main benefit is that there is no need to capture an animal to obtain a DNA sample. It can be particularly useful when studying endangered animals or those that live in a vast expanse, such as the open ocean. Other researchers have used the technique to detect the presence of organisms in a given area, such as freshwater fish in pond. This latest study marks the first time the technique has been used to provide information about an entire species. By learning more about such creatures as the endangered whale shark, researchers are hoping to find information that might help in preventing them from going extinct.
More information: Eva Egelyng Sigsgaard et al. Population characteristics of a large whale shark aggregation inferred from seawater environmental DNA, Nature Ecology & Evolution (2016). DOI: 10.1038/s41559-016-0004
Abstract
Population genetics is essential for understanding and managing marine ecosystems, but sampling remains challenging. We demonstrate that high-throughput sequencing of seawater environmental DNA can provide useful estimates of genetic diversity in a whale shark (Rhincodon typus) aggregation. We recover similar mitochondrial haplotype frequencies in seawater compared to tissue samples, reliably placing the studied aggregation in a global genetic context and expanding the applications of environmental DNA to encompass population genetics of aquatic organisms.
© 2016 Phys.org