Molecular 'sands of time' identified for fruitfly development
Okayama University have identified genes and processes responsible for pupation timing in the development of fruit fly larvae into adult insects.
The timing mechanism between specific developmental stages such as pupation in insects is so far little understood. In the Drosophila melanogaster—or common fruit fly—it is known that a pulse of the molting hormone ecdysone (E) is released to initiate the transition of the larva into a "prepupa" and a second pulse 10 hours later is then thought to induce pupation at around 12 hours. Using knock down models and forced expression of normal or modified proteins Hiroshi Ueda and colleagues at Okayama University investigated the role of different proteins on the timing of this 12 hour interval.
Shade, a protein mainly expressed in an insect organ that stores fat and is also known to sense nutrition—"fat body"—converts the hormone E into its active form 20-hydroxyecdysone (20E). Ueda and colleagues found that knock down of the shade gene in fat body delayed pupation whereas knockdown of the shade gene in other organs had no significant effect, confirming a role of fat body in pupation timing. "We think that the fat body may incorporate the nutritional status of the animal and sends a cue for the final decision of pupation,"say Ueda and colleagues in their report.
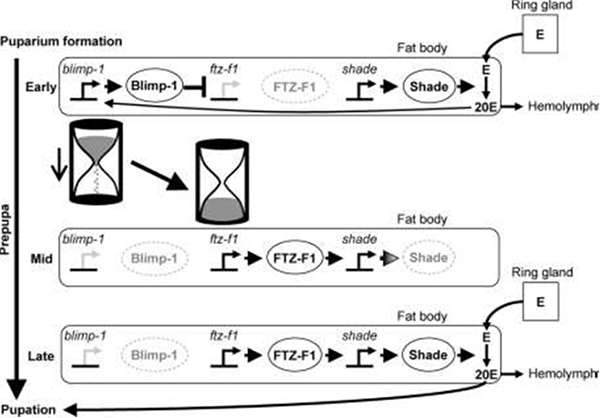
As for other influencers of pupation timing, the orphan nuclear receptor βFTZ-F1 - to which is attached another protein factor Blimp-1 - is known to be expressed after the 20E pulse. The researchers showed that over- and underexpression of βFTZ-F1 advanced and delayed pupation respectively. They also confirmed Blimp-1's role as a βFTZ-F1 repressor, showing that overexpression of Blimp-1 delayed pupation.
Blimp-1 is soon degraded—a protein conserved in mammals too. Studies with a line of Blimp-1 modified to enhance its stability led to delayed pupation and of the reduction of Blimp-1 expression led to advanced pupation. The researchers describe the naturally occurring Blimp-1 as like "sands in an hourglass." As the Blimp-1 molecules degrade and levels fall, βFTZ-F1 levels increase to initiate pupation. They also showed the advantage of repressor molecule such as Blimp-1 to use precise time measuring.
More information: Kazutaka Akagi et al. A biological timer in the fat body comprising Blimp-1, βFtz-f1 and Shade regulates pupation timing in, Development (2016). DOI: 10.1242/dev.133595
Journal information: Development
Provided by Okayama University