November 21, 2016 report
Molecular level investigation of water's hydrogen bonding network in salt solutions
(Phys.org)—What happens at the molecular level when salt dissolves in water? In introductory chemistry classes, we learn about the solubility of salts that dissociate into ions in water. Water has a complex hydrogen bonding network that should be disrupted by the presence of ions. However, studies have shown that there is little disruption to the hydrogen-bonding network. There is still debate as to the extent of molecular-level distortions that solvation of ions causes in water.
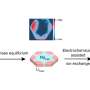
A group of researchers from the University of Zurich have elucidated the way ion solvation affects the hydrogen bonding network in water using a highly sensitive technique called 2D-Raman-terahertz spectroscopy. Their research shows a spectroscopic feature, called an echo, which was most pronounced in the dicationic chloride salts that they tested. This provides compelling evidence that the cations can structure the hydrogen bonding network. Their work appears in Nature Chemistry.
While the exact nature of the molecular effects of solvation was largely unknown, there have been certain macroscopic observations, including the viscosity of water increasing as ion charge density increases. One microscopic observation was that ions interact with water molecules near them (i.e., solvation), but it is unclear if these ions affect the hydrogen bonding network beyond the first solvation shell.
To investigate the molecular properties of solvation, Shalit et al. turned to 2D-Raman-terahertz spectroscopy. 2D-Infrared spectroscopy is helpful for detecting changes at short timescales; however this technique typically looks at the O-H peak, which is not sensitive enough to provide information on subtle changes on the hydrogen bonding network. The low-frequency regions (below 1,000 cm-1), however, can be used to detect changes in the intermolecular degrees of freedom in water.
2D-Raman-THz spectroscopy looks at the lower frequency vibrational regions of molecules and can be used to investigate vibrational signatures of the hydrogen-bonding network in water. It is sensitive enough to find subtle distinctions in the hydrogen bonding structure and can detect dynamic changes at femtosecond time scale.
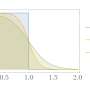
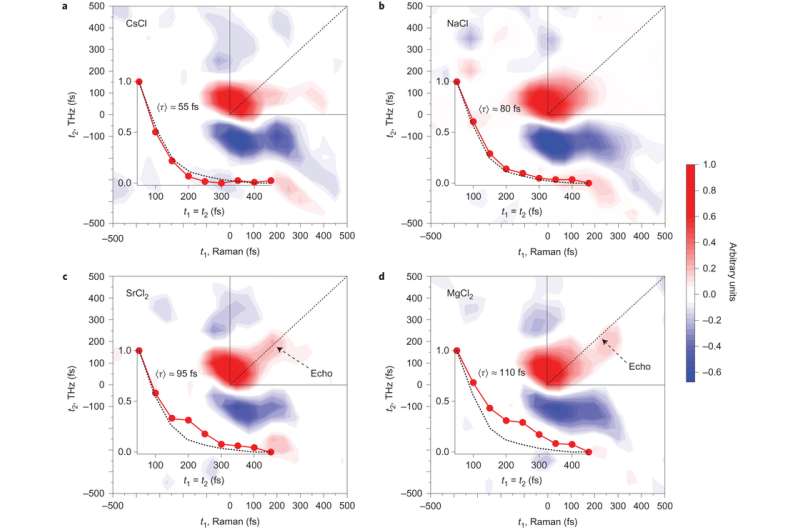
Recent studies on 2D Raman-THz spectroscopy of neat water showed that by conducting two perturbations, separated by a time t1, and reading out the response of the samples at time t2, can provide information on the degree of inhomogeneity in the hydrogen bond network. Inhomogeneity can be determined by the amount of signal along the t1=t2 line. The signal along the t1=t2 line, called an echo, indicates the system's ability to rephase after the second perturbation.
The current study uses this technique to look at how the chloride salts of monatomic cations Na+, Cs+, Sr2+, Mg2+ change the structure of water. Cs+ is considered a "structure breaking" ion and Mg2+ is considered a "structure making" ion. These are terms that reference the Jones-Dole equation for how the viscosity of water changes when ions are dissolved.
Shalit et al. showed that Cs+ looked the most like neat water based on the t1=t2 line. As the "structure making" ability of the cation increased, there was a longer relaxation time along the t1=t2 diagonal. This indicates greater inhomogeneity in the hydrogen bonding network. Sr2+ and Mg2+ showed the greatest deviation from neat water. Of the cations tested, these had the highest charge densities.
Overall, inhomogeneity of the hydrogen bonding network increased with increasing viscosity along the series Cs+ < Na+ < Sr2+ < Mg2+. For each of these the echo signature becomes progressively longer. Furthermore, additional analysis provided compelling evidence that the echo signal is not from water-ion vibrations, but from water-water vibrations. This means that the data was indicative of changes in the hydrogen bonding network, not ion-water interactions.
This study provides empirical data that correlates the macroscopic concept of viscosity when ions are dissolved in water with the microscopic heterogeneity of the hydrogen bonding network.
More information: Andrey Shalit et al. Terahertz echoes reveal the inhomogeneity of aqueous salt solutions, Nature Chemistry (2016). DOI: 10.1038/nchem.2642
Abstract
The structural and dynamical properties of water are known to be affected by ion solvation. However, a consistent molecular picture that describes how and to what extent ions perturb the water structure is still missing. Here we apply 2D Raman–terahertz spectroscopy to investigate the impact of monatomic cations on the relaxation dynamics of the hydrogen-bond network in aqueous salt solutions. The inherent ability of multidimensional spectroscopy to deconvolute heterogeneous relaxation dynamics is used to reveal the correlation between the inhomogeneity of the collective intermolecular hydrogen-bond modes and the viscosity of a salt solution. Specifically, we demonstrate that the relaxation time along the echo direction t1 = t2 correlates with the capability of a given cation to 'structure' water. Moreover, we provide evidence that the echo originates from the water–water modes, and not the water–cation modes, which implies that cations can structure the hydrogen-bond network to a certain extent.
Journal information: Nature Chemistry
© 2016 Phys.org