Scientists unravel weapons of defense against 'cotton cancer'
As world's largest cotton producer, China yields six to eight million tons cotton (30% of total world production) every year. However, high quality cotton cultivars are vulnerable to Verticillium wilt disease. Due to their long-term survival and vascular colonization of pathogen, the disease cannot be controlled by fungicides. How V. dahliae infects cotton was largely unknown and very limited resistance genes can be used for cotton breeding.
A group of Chinese scientists led by Prof. GUO Huishan from Institute of Microbiology of Chinese Academy of Sciences discovered that trans-kingdom small RNAs (sRNAs) can be used to protect crops from infection of fungal pathogens. This RNAi-based mechanism has been successfully manipulated to protect cotton from infection of a soil-borne fungal pathogen Verticillium dahlia (V. dahliae), the causal agent of vascular wilt disease so-called "Cotton Cancer" in China. This study has been published online in Nature Plants.
GUO's focus on the research of cotton-V. dahliae's infection process can be traced back to her visit in 2008 to Xinjiang in Northwest China, where she was astonished by the scene of huge area of infected cottons and desperate farmers. She began to try to search for a novel strategy to target V. dahliae.
Eight years of laboratory experiment and field research have brought GUO's team consecutive achievements in understanding Verticillium wilt disease.
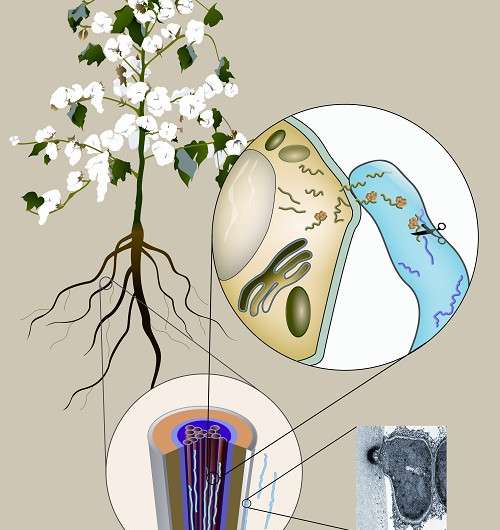
The team finds that trans-kingdom sRNAs exist during cotton-V. dahliae interactions, and some target virulence genes of pathogens. A novel penetration structure "hyphopodium" helps V. dahliae to "break" plant cell, which determines successful fungal infection (http://dx.doi.org/10.1371/journal.ppat.1005793). Most importantly, cotton cultivars expressing specific sRNAs are resistant to Verticillium wilt disease (http://dx.doi.org/10.1016/j.molp.2016.02.008).
Taking advantage of RNA and genome sequencing, the team found that cotton sRNAs exist in V. dahliae hyphae retrieved from infected cotton, which means sRNAs naturally transmit from host to pathogens during cotton-V. dahliae interaction.
They also demonstrated that some transmitted sRNAs target fungal genes were involved in pathogenicity and efficiently degraded gene transcripts. This phenomenon gave confidence to GUO that cotton would gain disease resistance by stably expressing sRNAs that target V. dahliae virulence genes.
"It was really time-consuming to generate the cottons stably expressing specific sRNAs, but we obtained them finally," said GUO. Their research proved to be successful both in the lab and in the field. sRNA expressing cottons showed 22.25% more disease resistance compared to the other cotton cultivars in the same cotton production base. The new generated cottons show resistance to wilt disease, especially in the field where the fungal populations are diverse.
More efficient fungal targets need to be found based on the established systems, which can also be guidelines for other un-controlled crop diseases in the future.
More information: Yun-Long Zhao et al, Hyphopodium-Specific VdNoxB/VdPls1-Dependent ROS-Ca2+ Signaling Is Required for Plant Infection by Verticillium dahliae, PLOS Pathogens (2016). DOI: 10.1371/journal.ppat.1005793
Tao Zhang et al. Host-Induced Gene Silencing of the Target Gene in Fungal Cells Confers Effective Resistance to the Cotton Wilt Disease Pathogen Verticillium dahliae, Molecular Plant (2016). DOI: 10.1016/j.molp.2016.02.008
Journal information: Nature Plants , PLoS Pathogens , Molecular Plant
Provided by Chinese Academy of Sciences



















