Researchers find way to ID many pathogens with few DNA probes

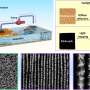
Rice University scientists have invented a technology that could potentially identify hundreds of bacterial pathogens simply, quickly and at low cost using a single set of random DNA probes. Rice's "universal microbial diagnostic," or UMD, uses pieces of randomly assembled DNA and mathematical techniques that were originally pioneered for signal processors inside digital phones and cameras.
In a paper online this week in Science Advances, Rice's research team used lab tests to verify that UMD could identify 11 known strains of bacteria using the same five random DNA probes. Because the probes are not specific to a particular disease, the technology provides a genomic-based bacterial identification system that does not require a one-to-one ratio of DNA probes to pathogenic species.
"If a laboratory today wants to test for 200 known pathogenic species, they need 200 different tests, each with its own specific DNA probe that was designed specifically to bind with DNA from a particular pathogen," said study co-author Richard Baraniuk, the lead scientist on the new study. "Our technology is fundamentally different. With a small set of DNA probes, we can test for a large number of species."
The new study includes several computer simulations, including one that shows how a random selection of five probes can identify 40 different strains of bacteria, and another that demonstrates how the system can accurately differentiate between 24 different species of Staphylococcus.
Baraniuk said UMD could help in treating and limiting the spread of antibiotic-resistant bacteria, which cause at least 2 million infections and 23,000 deaths each year in the United States, according to the Centers for Disease Control and Prevention.
"In many U.S. hospitals, it still takes several days to definitively identify the specific bacterium that's making someone sick," said Baraniuk, Rice's Victor E. Cameron Professor of Electrical and Computer Engineering. "The lack of rapid bacterial diagnostics can promote antibiotic resistance. Having an accurate, efficient and rapid system for identifying infectious pathogens quickly and inexpensively would help, and such a system would also be a valuable tool for public health, defense, global health and environmental science."
Historically, scientists identified bacteria by first culturing a sample—a process that takes several days—and then examining the organism under a microscope. More recent genomic identification methods that use polymerase chain reaction, or PCR, and genomic sequencing can be faster but require expensive equipment and training as well as a specific DNA probe for each pathogen to be tested. The probes are snippets of complementary DNA from a known disease. If the DNA from a patient's sample binds with the complementary DNA in a disease probe, the diagnosis is positive for that disease.
"Probe A is only good for finding bacterium A, and probe B is only good for finding bacterium B," said study lead author Amirali Aghazadeh, a graduate student in electrical and computer engineering in Baraniuk's lab. "Manufacturing such a probe for a new disease can take days to months and also requires expensive facilities that are only available in developed countries.
"With universal microbial diagnostic, we won't need new probes or to change any other parts of the sensing hardware," he said. "For any newly discovered bacterial strain, we can just adapt the software a little bit, and then the same platform can identify the new bacterium like any other."
Rather than identifying a target strain based on a 100 percent match with a specific probe, Rice's system tests how well the target DNA binds with several different random segments of complementary DNA. UMD uses a mathematical technique called compressive sensing, which was pioneered in the field of digital signal processing. With compressive sensing, the disease DNA need not bind with 100 percent of the probes. Instead, the UMD system measures how well the disease DNA binds with each of the random probes and creates a specific binding profile for the test organism. It then uses deductive reasoning to determine whether that profile matches the profile of any known pathogens.
"We believe the system will be most useful for rapidly and accurately looking for known targets, such as any organism on the World Health Organization's list of known pathogens, but another advantage of this method is that we get useful information even for organisms that have never been sequenced before," said study co-author Rebekah Drezek, professor of bioengineering and of electrical and computer engineering. "We'd be able to tell what pathogen a new disease is most closely related to."
More information: A. Aghazadeh et al. Universal microbial diagnostics using random DNA probes, Science Advances (2016). DOI: 10.1126/sciadv.1600025
Journal information: Science Advances
Provided by Rice University