Researchers generate whole-genome map of fruit fly genetic recombination
As eggs and sperm, or gametes, are formed during meiosis, chromosomes carrying the genetic material from each parent must find their partners, pair, and exchange parts of their DNA. This recombination is an important driving force behind genetic variability and evolution, but most importantly, it ensures that chromosomes move properly during the subsequent divisions that form these gametes. When recombination doesn't go smoothly, it can cause a number of problems in humans, including miscarriages and birth defects.
For the first time, researchers at the Stowers Institute have mapped where recombination occurs across the whole genome of the fruit fly Drosophila melanogaster after a single round of meiosis. Their results indicate that separate mechanisms position the two main kinds of recombination events, crossovers and non-crossovers.
The findings, which are reported online ahead of print in the journal Genetics, give important insights into the understanding of chromosomes and the mechanisms of inheritance.
"It is amazing to me that more than 100 years after the discovery of genetic recombination in flies, we are only starting to understand just how these events are distributed," says R. Scott Hawley, Ph.D., an investigator at the Stowers Institute and senior author of the study.
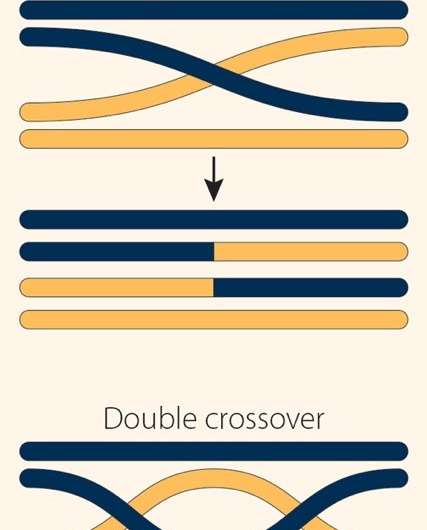
This genetic recombination takes place during a specialized form of cell division called meiosis. During meiosis, the cell copies all its chromosomes, pairs them up, and shuffles sections of genetic material between the arms of the paired or homologous chromosomes. This shuffling can occur one of two ways. In crossover events, large tracts of DNA are exchanged, like two people swapping playing cards. In non-crossover events, smaller pieces of DNA are copied from one arm and pasted onto another, like the crown from the King of Hearts in one player's hand suddenly appearing atop another player's King of Spades.
Once the deck is sufficiently shuffled, the cell divides, and then divides again to create four cells, each carrying only one copy of the organism's genome. Recombination ensures that each gamete ends up with a unique copy of every chromosome, but when the process goes awry, it can result in chromosomal abnormalities. For example, improperly placed crossovers or the complete lack of crossovers on chromosome 21 are a major cause of trisomy 21 or Down Syndrome in humans.
With one exception, all previous genetic studies of recombination in Drosophila have focused either on a single chromosome arm or on groups of flies pooled together. In this study, the Stowers researchers wanted to determine how both crossovers and non-crossovers are distributed across all five major chromosomal arms of fruit flies. Crossovers are relatively easy to identify because they involve entire chromosomal arms or parts of arms encompassing thousands of base pairs, the A's, C's, T's, and G's that make up DNA. But non-crossovers are tougher to spot, because they only involve a few hundred of those letters.
Therefore, Danny Miller, an MD-PhD student at the University of Kansas Medical Center who is conducting his doctoral research in the Hawley lab, had to rely on whole genome sequencing and new computer algorithms to pinpoint the locations of both kinds of events. Miller, lead author of the study, says the project gave him the opportunity to delve into the genetic principles that underlie human health and disease. "I would like to keep doing research as a physician-scientist when I graduate," says Miller, who plans to pursue a specialty in pediatric oncology or pediatric medical genetics. "I want to have a good foundational understanding of genetics. Some people may gloss over these basic questions, instead of looking at the data and trying to answer them."
In this study alone, Miller generated a vast amount of data. First, he mated two genetically distinct varieties of fruit flies, known to differ at roughly 500,000 different spots in their genetic code. Miller then sequenced the entire genomes of the resulting 196 progeny and wrote a custom computer program that could scan the 160 million bases of each fruit fly genome for evidence of recombination.
The approach identified a total of 541 crossovers and 291 non-crossovers. Unlike crossovers, which are generally distributed over the distal two-thirds of the chromosome arms, the non-crossovers were spread uniformly among the five major chromosome arms. Non-crossovers formed in places where crossovers rarely do, such as near the knotty centromere that ties the arms of chromosomes together. And they popped up close together, in contrast to crossovers that respond to a phenomenon known as interference when they try to form near other crossovers.
The researchers discovered that the number of crossovers varied widely not just according to their position along the chromosome arm, but also from chromosome to chromosome. For example, they identified five double crossovers on one arm of chromosome 2—fewer than expected.
"The finding gives credence to a certain kind of lore that has circulated in the field—the idea that each of the chromosomal arms was behaving a little differently—but nobody really knew for sure because nobody had looked at each of the arms in the same experiment," says Hawley. "What we found was that each chromosome has its own rules for making sure it gets its crossovers exactly where it wants them."
In addition to crossovers and non-crossovers, the researchers also noticed evidence of duplications and deletions caused by the presence of transposons, a special class of genetic elements that can jump from one area in the genome to another. These transposable elements are a serious problem for the meiotic machinery because they can skew the pairing of homologous chromosomes, triggering duplications or deletions of genetic material that affect the viability of an organism. The study identified transposable elements in 1-2 percent of the genome, in line with previous reports.
Journal information: Genetics
Provided by Stowers Institute for Medical Research