Researchers visualize over 10,000 proteins of fruit flies

The human genome codes for more than 20,000 different proteins, however the molecular role for many of these proteins is not known. As most proteins are conserved from fly to humans, understanding the molecular role of a protein in flies can be the first step towards a therapy against a variety of human diseases that are often caused by aberrantly behaving proteins. A consortium of scientists from the Max Planck Institutes of Biochemistry in Martinsried and Molecular Cell Biology and Genetics in Dresden, and the National Centre for Biological Sciences (NCBS) in Bangalore have now reached a milestone towards understanding the function of these proteins by using the fruit fly.
The human body is built by many hundreds of different cell types; each one has a very particular function in the body. Red blood cells transport oxygen, nerve cells exchange signals and muscle cells generate mechanical forces. The majority of cellular functions is produced by the action of 20,000-25,000 proteins coded in the human genome.
Although sequencing and annotation of human genome were completed in 2004, to date the function of many thousands of these proteins is still mysterious. It is often unknown, which cell types produce which proteins, and particularly where these proteins are located within the cells. Are they in the nucleus or within membranous vesicles, are they within neuronal dendrites or synapses, or are they within the contractile machinery of muscles? Protein localisation is an important piece of information, as it is the first step towards identifying a molecular function for a protein.
The proteins of a fly
Unravelling the function of a protein is often started in simpler model organisms such as worms or flies. Like humans, fruit flies have muscles, neurons, oocytes, sperm and many other essential cells types. The fly genome contains about 13,000 protein coding genes, which are responsible for building and maintaining all fly organs. Importantly, many of these proteins are very similar to the human proteins, thus studying a protein in flies will teach us about its role in the human body.
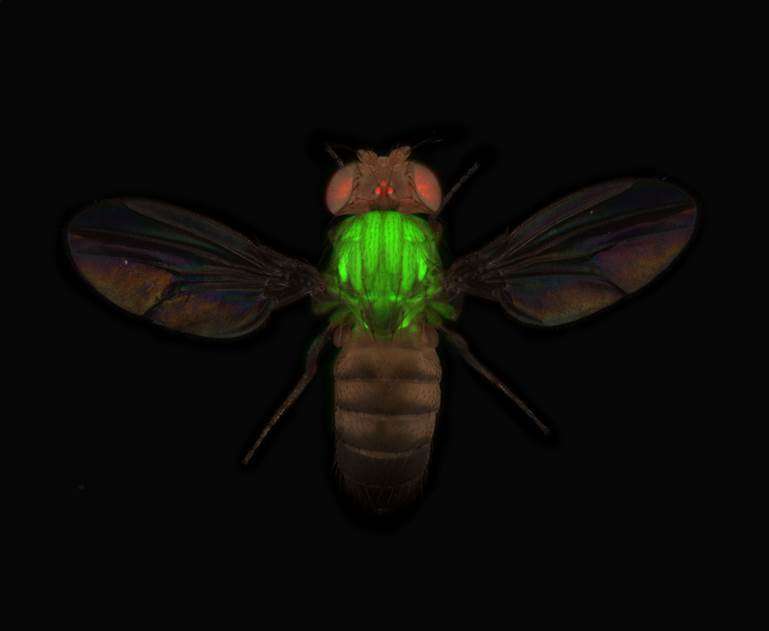
To boost these protein studies onto a systematic level, groups headed by Frank Schnorrer at the Max-Planck Institute in Martinsried, Pavel Tomancak and Mihail Sarov at the Max-Planck Institute in Dresden and K VijayRaghavan at the NCBS in Bangalore have generated a large resource for visualizing proteins in Drosophila melanogaster. By using modern molecular biology tricks, the scientists have attached a green fluorescent protein (GFP) tag to 10,000 of these protein coding genes in the test tube. Each tagged gene can then be re-introduced into the fly genome as a 'transgene', creating the fly 'TransgeneOme'. "Together, we thus far generated 880 different fly strains, each of which expresses a different fluorescently tagged protein", explains Frank Schnorrer, "these proteins can then be observed by fluorescent video microscopy in various cell types of the developing fruit fly".
For more than 200 proteins, the scientists documented where they are located during fly development, starting with an oocyte that develops into an embryo and finally into the mature fly. The Tomancak group used the so-called light sheet microscopy to film how proteins emerge in cells of the embryo during the first day of its development. The Schnorrer group used this resource to study the localisation of proteins in muscles. As in human skeletal muscles, fly muscles contain complex mini-machines called sarcomeres that produce the mechanical forces enabling animal movements.
"We have looked so far at only 200 of these transgenic lines. The future challenge lies in systematically imaging the localization of these proteins in many fly tissues and this is best achieved by involving the powerful Drosophila research community" predicts Pavel Tomancak. The resource will have enormous impact on the understanding of not only fly biology but also on the understanding of protein function in the different human cell types."
More information: Mihail Sarov et al. A genome-wide resource for the analysis of protein localisation in , eLife (2016). DOI: 10.7554/eLife.12068
Journal information: eLife
Provided by Max Planck Society