Molecular Lego with an encoded blueprint
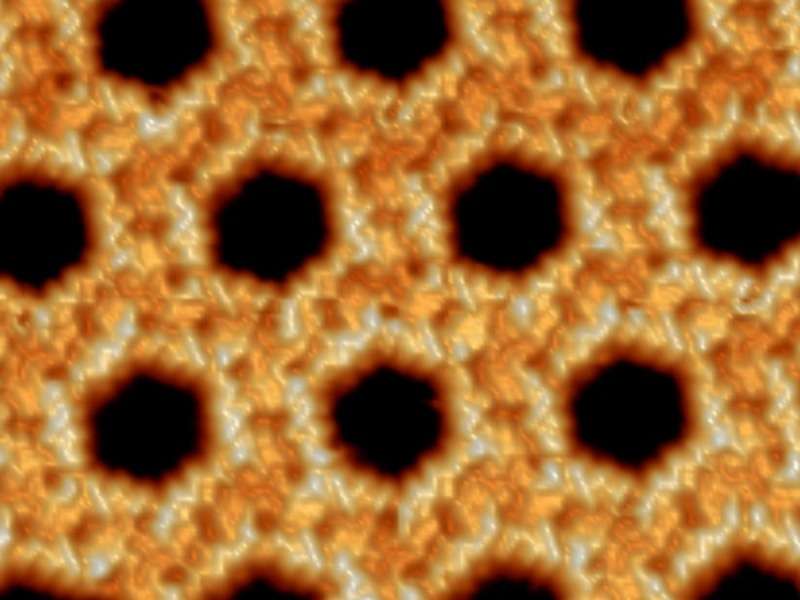

Nature contains a special kind of Lego brick: biological molecules, peptides to be exact, that can be built together to form a wide range of complex structures. Unlike the popular toy bricks, however, the molecular building blocks are flexible, can be stuck together using a number of different mechanisms and, under the right conditions, can even self-assemble using a blueprint encoded in each individual peptide. Scientists from the Max Planck Institute for Solid State Research have now succeeded in creating the conditions for self-assembly of a nano-scale honeycomb structure on a surface in the laboratory. A scanning tunneling microscope allowed them to observe the flawlessly arranged structure and see it in more detail than ever before. As peptides occur in countless variants, they can generate very different structures, some of which might be suitable for use as catalysts for chemical reactions.
Molecules are the building blocks of all life, just as Lego bricks are the building blocks of the world of play. This is why some researchers describe complex structures consisting of many molecules in terms of molecular architecture and compare nanotechnological formations with Lego structures. The analogy can be stretched even further, in that the studs on the top of a Lego brick fit into the gaps on the bottom of another in the same way as biological molecules bind to each other due to compatible forces between their atoms. Both Lego bricks and molecules can be separated again relatively easily, while the bricks or molecules themselves can only be broken using considerable force.
However, that is where the analogy between the molecular structures and toys ends, because in contrast to the rigid Lego bricks, molecules are flexible; and while there may well be several hundred different types of Lego bricks, the number of different molecules occurring in nature is far higher. Nature also boasts different sticking mechanisms: completely different atoms may exert forces between the molecules, and the forces themselves may be of different types. And finally, no tiny molecular hand is needed to put the assembly together; molecular assemblies build themselves. This self-organized process follows a construction plan that arises from the arrangement of their attachment sites.
Peptide composition and length determines structure
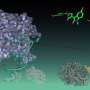
Scientists working with Klaus Kern, Director at the Max Planck Institute for Solid State Research, are now exploiting the properties of biological molecules to generate self-organized structures on surfaces. They place short peptides, angiotensin I and II to be precise, on a gold surface. Peptides are chain molecules and each link in the chain is an amino acid, of which 20 variants occur in nature. The specific structure generated on the surface depends on the composition and sequence of amino acids in the peptide in the same way that the functions of proteins, nature's molecular machines, depend on which amino acids they contain, in what sequence. The Stuttgart-based research team found that angiotensin I, which contains ten amino acids, forms two different network structures, both of which are relatively disordered and unstable. Angiotensin II, however, with its reduced sequence of just eight amino acids, forms a completely regular honeycomb structure with each wall consisting of two parallel peptides.
"This shows that sequence control, nature's tool for self-assembly, works on artificial surfaces too," says Stephan Rauschenbach, Research Group Leader at the Max Planck Institute in Stuttgart. This principle could be applied in nanotechnology to coat surfaces in order to give them certain properties. Nature already has countless catalysts at hand in the form of enzymes, most of which are also proteins, and their ability to accelerate chemical reactions could be transferred to surfaces; but even coatings with completely different functions, such as optical activity or electrical switching, might be possible, according to Stephan Rauschenbach.
High voltage to vaporize peptides
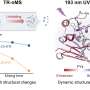
To deposit absolutely pure peptides onto the surface, the scientists use a technology called electrospray ion beam deposition. Although the neutral molecules cannot actually be evaporated, they can be coaxed into a gas phase by applying several thousands of volts of power to the solution. This charges the peptides and makes them repel each other so strongly that they prefer a gaseous state in order to keep their distance from one another. Since they are now electrically charged, the peptides in gaseous phase can be guided by electrical fields into an ultra-high-vacuum chamber. Free of any impurities, they are precipitated onto the gold surface in the chamber. Although initially distributed in a somewhat haphazard way, their thermal motion enables the molecules to come together at the binding sites.
No other method allows scientists to generate peptide structures in such pure form. Spotting a drop of peptide solution onto the surface achieves deposition but causes enormous contamination, and any attempt at evaporation just destroys the molecules. "It's like frying an egg," says Stephan Rauschenbach. "No matter how much you heat it, it will never evaporate. It just burns."
High-resolution imaging for precise modelling of structure formation
It was important to the Stuttgart-based researchers to generate the peptide assembly on the surface without impurities, primarily because this was the only way to observe the structure in near-atomic resolution using a scanning tunneling microscope. They cooled the sample to -230° Celsius, freezing even the molecular motion of the peptides. Using the high-resolution images, they then analyzed why each peptide's blueprint generated each resulting structure, as theoreticians used the detailed images to generate accurate models to explain the self-assembly mechanisms.
"As these peptides consist of large, unwieldy amino acids, they are fairly rigid, a bit like Lego bricks," explains Sabine Abb, an important contributor to the study. Another consideration is that peptides contain both polar and non-polar amino acids, and the assembly blueprint varies according to how they are distributed. Polar amino acids form strong bonds with their neighbouring peptides, while non-polar amino acids are more passive. This interplay of bonding and passivity is the main basis on which the peptides self-assemble into defined structures. Too many polar amino acids and the peptides stick together at random, so no real structure ensues; too few, and the structures formed are unstable.
"Now that we have a better understanding of how to apply nature's tool of self-organization on surfaces, we just have to find the right sequences for structures we want to produce," observes Stephan Rauschenbach. "It won't be easy," he cautions. "After all, it took nature billions of years to create enzymes with all their specific functions."
More information: Sabine Abb et al. Two-dimensional honeycomb network through sequence-controlled self-assembly of oligopeptides, Nature Communications (2016). DOI: 10.1038/ncomms10335
Journal information: Nature Communications
Provided by Max Planck Society