Research might lead to a cheaper, more efficient, hydrogen economy
Researchers have worked out how some bacteria rip hydrogen apart to produce energy, just like a biological fuel cell. In the process they question the current ideas of how this happens and possibly move one step closer to a cheaper, more efficient, hydrogen economy. The research is published in the journal Nature Chemical Biology.
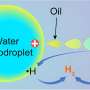
The scientists used a combination of different techniques to work out what happens at the heart of bacterial enzymes called nickel iron hydrogenase. One of the authors Professor Simon Phillips, Director of the Research Complex at Harwell in Oxfordshire said: "The hydrogenases can do two things with hydrogen gas, turn it into protons and electrons or recombine them to form hydrogen. This can be done in a fuel cell but with platinum. Evolution has produced bacteria that can do it with nickel and iron, cheap and plentiful metals." These bacteria use hydrogen to provide energy; the enzymes convert hydrogen into electrons, just like a hydrogen fuel cell where electricity is generated.
Hydrogen fuel cells are seen as a vital part of our low carbon future, using hydrogen to produce electricity to drive vehicles rather than fossil fuel burning internal combustion engines. However, fuel cells require rare and expensive platinum, this enzyme does the same trick with cheap and plentiful iron and nickel. What's more, this enzyme can do the reaction in reverse, producing hydrogen from electricity and water. As a result the study of these enzymes is a very active and important area of research.
Enzymes are nature's machines made of protein, and have evolved over millions of years to be very good at what they do, often better than the equivalent human technology. These hydrogenases use cheap metals to do at room temperature what fuel cells can only do with pricey platinum and at high temperatures. If the mechanism of this enzyme can be adapted then it could make far cheaper fuel cells, and also a way of making hydrogen cheaply from electricity and water – two highly sought after technological advances. That, though, is still a long way off.
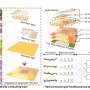
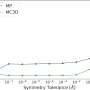
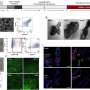
To explore how the hydrogenase works in detail the researchers made subtle alterations, tiny mutations, to each of the amino acids in the pocket in the enzyme where the hydrogen reaction takes place. This resulted in a version that appeared to be the same as the original but was far less efficient. The key here was that they had to know whether the changes had altered the shape of the enzyme or just its activity. To do this they used Diamond Light Source, the UK's synchrotron science facility, to work out and compare the structures of the original and changed enzyme using a technique called x-ray crystallography. Professor Phillips said: "We got very accurate structures of the mutant and basically nothing moved apart from the atoms of the amino acids. So the loss of activity is not down to a change in structure or the loss of the metals. Diamond was crucial in determining this. "
This confirmed that reduction in activity had to be due to chemical, not physical, changes. The tiny change removed a nitrogen atom at its heart, one that was essential to make the hydrogen reaction work.
What they discovered was that the enzyme uses a principle known as a Frustrated Lewis Pair. A normal Lewis pair is made up of different chemicals that are keen to interact with each other and would so given the opportunity. In this case these are the atoms of nickel and iron together, and a particular nitrogen atom built into the enzyme. The frustration bit is that in the enzyme, these entities are held close enough to see each other, but not close enough to interact fully. This produces an area of tension between them, a bit like holding a dog back from its food bowl. And just as anything getting between a dog and its food is at risk of being mangled, so a hydrogen molecule fed into this area of tension gets split apart.
The concept of a Frustrated Lewis Pair is relatively new even though nature has been successfully using it for likely billions of years. The mechanism is reasonably well understood but is different from the textbook explanation of how these enzymes work. Professor Phillips said: "Our research shows unequivocally that the chemical mechanism in the literature is wrong."
Now that this research has revealed just how the enzyme splits hydrogen the researchers are keen to go further and see if they can watch it in action.
One way they hope to do this is to use Diamond's latest innovation, a vacuum chamber for x-ray crystallography – the only one in the world to date. Normally x-ray crystallography is carried out in air, but the air molecules interfere with the x-rays and introduce noise to the experiment, a bit like static on a badly tuned radio. This can obscure the very weak signal given by the hydrogen inside the enzyme. By sucking out all the air the researchers can reduce this noise considerably and stand a good chance of actually spotting the crucial hydrogen and exactly what happens to it. Simon Phillips said: "This unique facility means that we might improve the signal to noise ratio by a factor of ten which could let us see the hydrogen in place."
More information: Rhiannon M Evans et al. Mechanism of hydrogen activation by [NiFe] hydrogenases, Nature Chemical Biology (2015). DOI: 10.1038/nchembio.1976
Journal information: Nature Chemical Biology
Provided by Diamond Light Source