December 3, 2015 report
Diagnostic tools using synthetic biology
(Phys.org)—Synthetic biology is a multi-disciplinary field that applies engineering techniques to biological systems. While the foundations of synthetic biology were laid in the 1990s with the burgeoning of genomics and automated sequencing, synthetic biology as a discipline, emerged in the early 2000s with the development of simple gene regulatory circuits. Since then, the field has grown beyond making simple circuits to developing diagnostic tools.
Shimyn Slomovic, Keith Pardee, and James J. Collins of MIT and Harvard discuss the current trends in synthetic biology to make diagnostic tools, and what additional hurdles must be overcome before these methods can be applied in the field or in the clinical setting. Their article is part of a special hundredth anniversary series in the Proceedings of the National Academy of Sciences commemorating exceptional research published in the journal over the last century.
The rational design of diagnostic tools in synthetic biology uses natural regulation and detection systems to design sensors. Ideally, these sensors are selective, sensitive, and provide some kind of output, such as fluorescence or a color change, when it has detected the target cell, environmental cue, or pathogen. Slomovic, Pardee, and Collins discuss three broad categories of synthetic biology-based diagnostic tools: whole-cell biosensing, in vitro diagnostics, and in vivo diagnostics.
Whole-cell biosensing
Microorganisms can be employed as a kind of biosensor. One well-known example is photosynthetic cyanobacteria incorporated into hardware. The cyanobacteria are redox active. Electron transfer through the hardware will produce a current, but herbicides quench its redox activity. The lack of current serves as an indicator for the presence of herbicides in places such as natural water reservoirs.
More recent research involves tailoring microorganisms using genetic engineering. Engineering methods allow scientists to design bacteria that are selective for a particular substance. For example, bacteria have been engineered to detect arsenic in water. These genetically engineered bacterial cells have been integrated into circuits and other mechanical components to produce a hybrid device. However, hybrid devices have lost popularity, giving way to plate readers which have a higher throughput.
RNA-based biosensing can detect metabolites or RNA sequences that are specific to a particular disease. RNA biosensing works by releasing stalled RNA translation using a synthetic piece of RNA that contains a complementary portion of the target sequence. One of the difficulties with this technique, however, is that the RNA sensing portions are confined within the bacterial cell and are not accessible to the external environment.
In vitro diagnostics
Bacteriophages are nature's bacteria sensors. They are viruses that home in on particular bacteria and infect them. Engineered bacteriophages are relatively inexpensive to make and can be designed to provide output indicators, such as bioluminescence, when it has detected the presence of a particular bacterial strain. Bacteriophages have been used both as sensors and as alternatives to antibiotics. For example, phages allow scientists to detect the presence of bacteria associated with the bubonic plague in a human blood sample in a matter of hours.
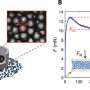
Paper-based diagnostics are a cheap, mobile method that shows promise for eventual field use. Using commercially available bacterial or mammalian transcription/translation systems, scientists are able to freeze-dry these systems on to paper or other porous materials. These systems target certain RNA sequences in a similar way to the RNA-based biosensing, mentioned above, but the circuitry is on paper rather than within the cell. The system becomes active when rehydrated, allowing the freeze-dried material to be easily transported. Recent tests with an Ebola virus disease detection system used color-changing enzymes for easy output.
In vivo diagnostics
Real-time biosensing is attractive for monitoring changes in the organism's internal environment. This can be done with engineered bacteria that are administered to the organism. One example is the use of engineered E. coli to monitor the gut microbiome. In mouse studies, E. coli. were engineered to detect and record exposure of the gut microbiome to a particular drug.
Mammalian cells can also function as in vivo biosensors, and they have the added benefit of possibly serving as both the sensor and the therapy. These cells utilize synthetic gene networks that can either change or monitor the organism's cellular environment. Additionally, cellular biosensing allows the synthetic circuit to be isolated from the environment but also able to interact with it.
One way that mammalian cells have been used for diagnostics is to monitor the expression profiles of certain cells. Cells express, or "turn on", some genes but not others. The particular set of genes that are turned on provides an expression profile. Some cancer cells, cells that have been infected by a virus, or cells whose immune system is malfunctioning (autoimmune) have different expression profiles from normal cells. These cells can be targeted and identified using engineered mammalian cells. This has the added benefit of potentially destroying the diseased cell either by turning on the cell's self-destruct genes or by drug targeting.
Unfortunately, the biggest difficulty with in vivo diagnostics is delivering the engineered cell or synthetic genetic construct to the right place. Slomovic, Pardee, and Collins point out that this is where the field of nanobiotechnology comes into play. Recent research shows that some nanoparticles will home in on certain cells types, and others have been looking at lipid-based vesicles as another possibility for targeted delivery.
Slomovic, Pardee, and Collins point out that "success in synthetic biology will continue to be found at the interfaces between disciplines." Synthetic biology, while still a relatively new field, draws from years of work in genetics, microbiology, engineering, and nanotechnology, and by combining these fields, researchers have been able to make formidable progress in biosensing and in vitro and in vivo diagnostics.
More information: Shimyn Slomovic et al. Synthetic biology devices for in vitro and in vivo diagnostics, Proceedings of the National Academy of Sciences (2015). DOI: 10.1073/pnas.1508521112
Abstract
There is a growing need to enhance our capabilities in medical and environmental diagnostics. Synthetic biologists have begun to focus their biomolecular engineering approaches toward this goal, offering promising results that could lead to the development of new classes of inexpensive, rapidly deployable diagnostics. Many conventional diagnostics rely on antibody-based platforms that, although exquisitely sensitive, are slow and costly to generate and cannot readily confront rapidly emerging pathogens or be applied to orphan diseases. Synthetic biology, with its rational and short design-to-production cycles, has the potential to overcome many of these limitations. Synthetic biology devices, such as engineered gene circuits, bring new capabilities to molecular diagnostics, expanding the molecular detection palette, creating dynamic sensors, and untethering reactions from laboratory equipment. The field is also beginning to move toward in vivo diagnostics, which could provide near real-time surveillance of multiple pathological conditions. Here, we describe current efforts in synthetic biology, focusing on the translation of promising technologies into pragmatic diagnostic tools and platforms.
Journal information: Proceedings of the National Academy of Sciences
© 2015 Phys.org