Innovative imaging technique reveals new cellular secrets

Cellular mitosis depends in part on small organelles that extend spindles to pull apart chromosome pairs. Before they can perform this and other essential tasks, these tiny cylindrical structures—known as centrioles in animals and spindle pole bodies (SPBs) in yeast—must themselves duplicate.
However, much about this nanoscale process has remained veiled by the limits of current microscopy. Optical approaches cannot resolve objects below certain wavelength limits, while non-optical approaches like electron microscopy (EM) can only study nonliving cells.
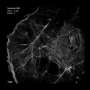
Now, a team of researchers from the Stowers Institute for Medical Research and the University of Colorado Boulder has devised a novel optical technique—a combination of structured illumination microscopy (SIM) and single-particle averaging (SPA)—to resolve individual components of SPB duplication in living yeast cells. In the process, they have uncovered surprising facts in what many once considered well-trodden ground. More than that, they have opened up new possibilities in the field of cellular imaging.
"The use of SIM to study SPB structure completely changes the types of questions we can ask and answer, since sample size is no longer limiting and it is likely that SIM will work in living cells," says Sue Jaspersen, Ph.D., an associate investigator at the Institute who led the investigation.
The study will be published in the online journal eLife on September 15, 2015.
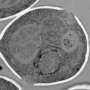
For years, EM, which uses a beam of electrons to achieve molecular and even atomic resolutions, has been the go-to technique for studying SPBs, which at less than 200 nanometers (nm) in size fall below the wavelength limit of what is observable using visible light. However, EM carries with it significant limits, including the fact that it does not work on living cells.
The research team turned to SIM as an optical alternative. SIM uses a laser-generated field of horizontal lines to project an interference pattern onto a sample. According to Jay Unruh, Ph.D., a Stowers research advisor and co-author, analyzing these patterns enables researchers to effectively double their resolution.
"Basically, finely structured objects interfere with the line pattern in a way that makes a new pattern, which is larger in size."
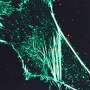
For all of its advantages, SIM still involves sifting through a great deal of noise. To deal with this problem, and to better localize the subjects under study—which can assume various shapes and positions—the team turned to SPA. In this technique, researchers align a large number of images along reference points in three-dimensional space and then average them into a single, characteristic image. The result is a sharper, more reliable picture of what is going on within the cell.
"Small deviations that can be due to noise are eliminated, and we can localize proteins with high confidence, often with even greater precision than via SIM alone," says Jaspersen. "We estimate the precision to be in the 10-30 nm resolution range."
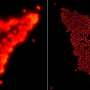
The SPA-SIM technique made up part of a two-color structured illumination microscopy approach that used endogenously expressed fluorescent protein derivatives. Through genetic techniques, the researchers fused the proteins in question to two standard fluorescent proteins, which would show up when the yeast naturally expressed the gene.
According to Unruh, this study represents the first combined use of SPA with SIM, and one of the first dual-color super-resolution SPA papers. Moreover, where other studies concentrated on a few protein pairs, the current study "characterized a significant portion of a large, many-protein complex."
"The big-picture take away is that even structures that are considered by many to be 'solved'—we know what they look like by EM, we know all the parts, we know many of the physical interactions between those parts—have remarkable surprises when one is finally able to study their formation in cells."
Among those surprises: SPB duplication, once thought to occur in the G1 period of interphase, now appears to begin near the end of mitosis. Moreover, the structures by which SPBs attach to the nuclear envelope do not form at the end of duplication, as once thought, but rather during the duplication process itself.
The study also reported a number of never-before-seen structural features, including "the structure and timing of half-bridge elongation, the composition of the satellite and the formation of the membrane pore."
But that's just the beginning. According to Jaspersen, the SPA-SIM technique is applicable to a wide variety of subjects beyond SPB structure.
"This method can be applied to any regular intracellular structure if there is some known reference protein that can be used."
Other Stowers authors are Shannon Burns, Zulin Yu, Ph.D., Sarah Smith, Ph.D., and Brian Slaughter, Ph.D. Additional contributors include Jennifer Avena, Ph.D., and Mark Winey, Ph.D., at the University of Colorado, Boulder.
The study was funded by the Stowers Institute and the National Institutes of Health (Mark Winey, P01 GM105537; Jennifer Avena, 5 T32 GM007135; Jennifer Avena, Mark Winey, R01 GM51312). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Lay Summary of Findings
Key parts of cell division rely on tiny, tube-shaped structures, called spindle pole bodies (SPBs) in yeast. While SPBs are well studied, many questions remain about how they copy themselves. Partly, this blind spot is due to the limits of optical microscopes, which can only see objects that are larger than the wavelengths of light with which they are viewed. A common alternative, electron microscopes, can see much smaller objects, but do not work on living cells.
In a study to be published on the eLife website on September 15, 2015, a team of researchers from the Stowers Institute for Medical Research and the University of Colorado Boulder combined two optical systems in a new way to get around the natural limits of optical microscopes. One, called structured illumination microscopy (SIM), makes laser-based interference patterns that change based on what they interact with, doubling the resolution of optical microscopes. The other, single-particle averaging (SPA), brings tiny objects and their locations into sharper focus by averaging many images into one "typical" picture. Using this method, the team found that SPBs duplicate and form some structures at different times than once thought. They also spotted a number of never-before-seen structures used in SPB duplication.
Journal information: eLife
Provided by Stowers Institute for Medical Research