How to preserve fleeting digital information with DNA for future generations

Hand-written letters and printed photos seem quaint in today's digital age. But there's one thing that traditional media have over hard drives: longevity. To address this modern shortcoming, scientists are turning to DNA to save unprecedented amounts of digital data for posterity. One team has demonstrated that DNA they encapsulated can preserve information for at least 2,000 years, and they're now working on a filing system to make it easier to navigate.
The researchers present their work today at the 250th National Meeting & Exposition of the American Chemical Society (ACS).
"If you go back to medieval times in Europe, we had monks writing in books to transmit information for the future, and some of those books still exist," says Robert Grass, Ph.D. "Now, we save information on hard drives, which wear out in a few decades."
At the same time, digital technology has spurred an explosion in the amount of information available at any given moment. Any new techniques scientists develop to preserve even parts of our digital universe would have to be extremely small. This is where DNA comes in.
"A little after the discovery of the double helix architecture of DNA, people figured out that the coding language of nature is very similar to the binary language we use in computers," says Grass, who is with ETH Zurich. "On a hard drive, we use 0s and 1s to represent data, and in DNA, we have four nucleotides A, C, T and G."
But DNA has two major advantages over hard drives: size and durability. An external hard drive about the size of a paperback book can back up five terabytes of information and might last 50 years. In theory, a fraction of an ounce of DNA could store more than 300,000 terabytes. And, from archaeological finds, scientists know that DNA from hundreds of thousands of years ago can still be sequenced today.
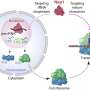
A handful of research groups are exploring methods to take advantage of DNA's storage potential. Grass' team has encoded DNA with 83 kilobytes of text from the Swiss Federal Charter from 1291 and the Method of Archimedes from the 10th century. They encapsulated the DNA in silica spheres and warmed it to nearly 160 degrees Fahrenheit for one week, which is the equivalent of keeping it for 2,000 years at about 50 degrees. When they decoded it, it was error-free.
Now that the researchers have demonstrated how to synthetically preserve DNA for long periods of time, they're tackling the next challenge.
"In DNA storage, you have a drop of liquid containing floating molecules encoded with information," Grass says. "Right now, we can read everything that's in that drop. But I can't point to a specific place within the drop and read only one file." So, he and his colleagues are currently developing ways to label specific pieces of information on DNA strands to make them searchable.
Like many technologies in their early years, DNA storage comes with a hefty price tag. Encoding and saving a few megabytes of data costs thousands of dollars, Grass says. In other words, consumers won't have the option of buying DNA-based hard drives anytime soon.
So what will this technology accomplish? Grass says that question has yet to be answered. If it were up to him, he says he would take data snapshots of the ever-evolving Wikipedia, for example, to preserve its various iterations so they're not lost forever as users make edits. DNA storage also could preserve troves of historical texts, government documents or entire archives of private companies, all in a droplet.
"This interest in preserving information is something we have lost, especially in a digital world," he says. "And that's what I'd like to help address and encourage people to do: Save information we have today for future times."
More information: The engineering of DNA for the long-term storage of digital information, the 250th National Meeting & Exposition of the American Chemical Society (ACS).
Provided by American Chemical Society