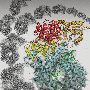Researchers construct atomic model of an immature retrovirus
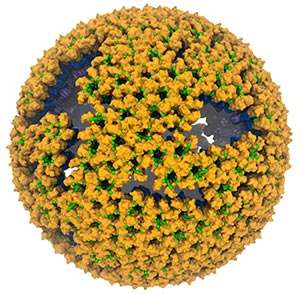
Using molecular modeling and large-scale molecular dynamic simulation, University of Illinois researchers constructed an atomic model of an immature retrovirus.
The researchers, from the Theoretical and Computational Biophysics Group at the Beckman Institute for Advanced Science and Technology at Illinois, published their work in the journal Structure.
Retroviruses, such as HIV, are tricky to treat. They go through a multistage process to produce infectious particles. The viruses that are released from infected cells are initially in an immature state and are composed of an RNA genome surrounded by a coat of protein. Upon their release, the viruses undergo a maturation process that rearranges the viral proteins and activates the conversion of the RNA genome into DNA through a process called reverse transcription. The viral DNA then invades the host cell's genome. The infected cell is programmed to release multiple copies of the immature virus into the host's bloodstream. These newly released viruses must in turn mature before they can infect other cells.
Not only do retroviruses cause life-long infections, but also in the complex process of reverse-transcribing RNA into DNA, many mutations can occur, making the retrovirus even more difficult to target.
One strategy for preventing the spread of a retrovirus is to lock the viral particles in their immature, non-infectious state. Unfortunately, the complexity and size of the viral particle—an irregular and incomplete hexagonal shell of close to 100 nanometers in size—have prevented the experimental determination of the atomic-level structure of the particle.
A number of studies have examined the structure of the Rous sarcoma virus (RSV), which affects birds and provides a good model for other retroviruses, but none have been able to provide a high-resolution look at the immature stage of the virus.
Over a period of two years, researchers have been performing calculations and simulations in order to reveal the structural features of the virus. With the help of the Titan supercomputer at the Oak Ridge Leadership Computing Facility in Tennessee and the Blue Waters supercomputer at the National Center for Supercomputing Applications (NCSA) at Illinois, the researchers were the first to provide the atomic-level structural model of the immature retroviral lattice of RSV. A five-microsecond simulation describing the motion of half a million atoms involves a hefty calculation that took five days using 30 percent of the Titan supercomputer, more than 5,000 computing nodes—each node is equivalent to a high-end workstation.
"We have had a pretty good understanding of the mature infectious particle at a level where we can make specific predictions about the local chemical interactions between the protein subunits in the virus," said Rebecca Craven, professor of microbiology and immunology and at Penn State University and one of the study's authors. "But the field was really lacking similar high-resolution knowledge about the immature virus. This new model is the first to give us an atomic-level look at the immature state. With that knowledge we can try to understand the precise molecular mechanisms of virus maturation and help to elucidate how drugs can be designed to interfere with that."
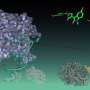
According to Boon Chong Goh, a graduate student in physics at Illinois and lead author on the study, a six-helix bundle domain located on the inside surface of the immature protein shell could be a key to understanding and blocking the virus.
"Recent advances in cryo-electron tomography allow us to model the majority of the immature retroviral lattice, except the six-helix bundle domain," said Goh. "Experimentalists do not have a clear view of that domain because of its high flexibility. And that's where we come in. Using advanced computational techniques and supercomputers, we modeled and refined an all-atom model of the six-helix bundle.
The researchers believe that the six-helix bundle is amphipathic, a chemical property that possesses an affinity for both water and fat, and that it is enclosed by a ring of salt bridges, contributing to the stability of the bundle.
"HIV is a close relative to RSV, and HIV is also known to have this domain, which has been a drug target for years. A drug named Bevirimat (BVM) was developed to target the six-helix bundle of immature HIV, but it did not pass the clinical trials," said Goh.
"The experience with BVM did show that inhibitors that target the six helix bundle domain can be very powerful HIV anti-retroviral drugs by preventing virus maturation," said Craven.
"The main idea is that you have two forms of the virus: the immature and the mature," said Juan Perilla, postdoctoral researcher and co-first author of the study. "The immature is not infectious, so the idea is that ultimately you want to prevent it from becoming the mature form. The problem is that BVM targets the six-helix bundle domain, but no one really knows the structure of the immature lattice in HIV. There are a few models, but they are not high resolution, so we decided to work in this direction and we picked RSV because it's a good model to study the virus. Our next step is to go to HIV."
"This result is an example of where computational methodology is really complementing medical lab experiments," said Klaus Schulten, director of the TCBG and professor of physics.
According to Schulten, computational experiments can help provide solutions to complex problems in the laboratory.
"The living world is made of molecules and you need to know this world. Sometimes you just do it through trial and error in a chemical research lab and you see that a newly designed drug molecule has a certain effect that often you only understand later. But then there is, of course, a more systematic way of resolving the molecular world, namely the computational way taken in our RSV study, and one can derive medical treatments from the knowledge reached computationally," Schulten explained. "This will be done more and more often."
The group has already begun the process of studying the immature HIV virus on the Blue Waters supercomputer located at the National Center for Supercomputing Applications at Illinois.
Journal information: Structure