April 23, 2015 report
Researchers develop computer model of bacterial colony interactions
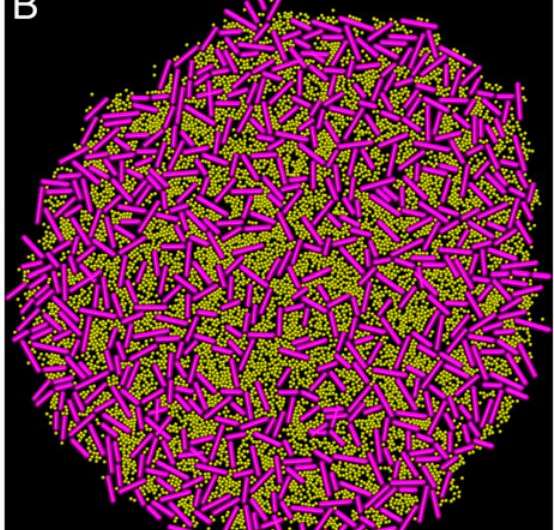
(Phys.org)—One of the remarkable properties of bacterial colonies is the self-ordering aggregation and orientation of bacterial cells. Bacteria secrete extracellular polymeric substances (EPS) that form biofilm-like structures in which the cells aggregate. Due to the density of such systems, researchers have speculated that the high degree of order manifested by colonies is due to specific mechanical interactions between the cells and the biofilm matrix.
A group of researchers from Rice University, Columbia University, and Tel Aviv University developed a computer model of bacterial cell-EPS interactions that allowed them to change variables and study the effects on the aggregation and orientation of bacterial cells. Published in The Proceedings of the National Academy of Sciences, their study confirms the influence of mechanical effects on the heterogeneity of cell distribution in bacterial colonies.
Sim MicroCity
At low densities, cells communicate primarily through chemical means. At higher densities, however, direct mechanical interaction between cells becomes a governing factor in self-organization. The researchers built a simple model examining the interactions of nonmotile, rod-shaped bacterial cells within a self-produced EPS to determine how a nonequilibrium bacterial colony undergoes spontaneous aggregation.
The particle-based model is restricted to two dimensions, thus omitting the overlapping of bacterial cells that occurs in reality. The surrounding polymeric EPS is approximated by small spherical particles, while bacteria are modeled as larger, growing spherocylinders with a constant diameter. This simplified computer model thus describes the aggregation characteristics of the bacteria via their x-y coordinates and unit vectors representing the orientation of their symmetry axes.
Mechanical interactions driven by repulsion forces
The study treats EPSs as depleting particles. Setting high depletion forces within the medium results in high repulsion forces between cells and the EPS. This interaction is implemented in the model by allowing the repulsion such that the centers of EPS particles are restricted from coming into an excluded volume zone surrounding the bacteria. The origin of the depletion force is an entropy gain that occurs when bacterial cells approach one another, thereby reducing the excluded volume for the EPS.
Among their findings, the researchers noted that an increase in the cell-EPS repulsive force resulted in considerable aggregation of cells and a high degree of self-organization within the colony. To confirm this effect of repulsive elastic force between the cells, the researchers increased the rate at which the cells secreted EPS, while keeping the repulsive forces the same. The result was a colony of almost heterogeneously distributed bacterial cells in a medium of EPS particles and very little segregation. They write, "It is thus evident that a strong repulsive mechanical interaction is needed to drive the system, leading to aggregation of bacterial cells by depletion attraction, even if there are excess depletants (EPS particles) in the medium."
For future studies, the researchers suggest that their model would be easily applied to a number of three-dimensional scenarios found in real life. Adding a third dimension would allow them to include the effects of buckling transitions that drive cells in dense colonies upward. The current model also omitted effects of cell death within a system. Self-driven, motile bacteria could be modeled with parameters similar to the current study. And the dynamics of cells and the extracellular matrix could also be modeled in animal tissue conditions.
More information: "Mechanically-driven phase separation in a growing bacterial colony." PNAS 2015 ; published ahead of print April 13, 2015, DOI: 10.1073/pnas.1504948112
Abstract
Secretion of extracellular polymeric substances (EPSs) by growing bacteria is an integral part of forming biofilm-like structures. In such dense systems, mechanical interactions among the structural components can be expected to significantly contribute to morphological properties. Here, we use a particle-based modeling approach to study the self-organization of nonmotile rod-shaped bacterial cells growing on a solid substrate in the presence of self-produced EPSs. In our simulation, all of the components interact mechanically via repulsive forces, occurring as the bacterial cells grow and divide (via consuming diffusing nutrient) and produce EPSs. Based on our simulation, we show that mechanical interactions control the collective behavior of the system. In particular, we find that the presence of nonadsorbing EPSs can lead to spontaneous aggregation of bacterial cells by a depletion attraction and thereby generates phase separated patterns in the nonequilibrium growing colony. Both repulsive interactions between cell and EPSs and the overall concentration of EPSs are important factors in the self-organization in a nonequilibrium growing colony. Furthermore, we investigate the interplay of mechanics with the nutrient diffusion and consumption by bacterial cells and observe that suppression of branch formation occurs due to EPSs compared with the case where no EPS is produced.
Journal information: Proceedings of the National Academy of Sciences
© 2015 Phys.org