Super-resolution microscopes reveal the link between genome packaging and cell pluripotency
A study using Super-resolution microscopy reveals that our genome is not regularly packaged and links these packaging differences to stem cell state. A multidisciplinary approach allowed scientists to view and even count, for the first time, the smallest units for packaging our genome. This study has brought together biologists and physicists from the Centre for Genomic Regulation and the Institute of Photonic Sciences, both in Barcelona.
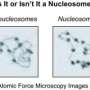
In 1953 Watson and Crick first published the discovery of the double helix structure of the DNA. They were able to visualize the DNA structure by means of X-Ray diffraction. Techniques, such as electron microscopy, allowed scientists to identify nucleosomes, the first and most basic level of chromosome organisation. Until now it was known that our DNA is packaged by regular repeating units of those nucleosomes throughout the genome giving rise to chromatin. However, due to the lack of suitable techniques and instruments, the chromatin organisation inside a cell nucleus could not be observed in a non-invasive way with the sufficient resolution. Now, for the first time, a group of scientists at the CRG and ICFO in Barcelona, have been able to visualise and even count the smallest units which, packaged together, form our genome. This study was possible thanks to the use of super-resolution microscopy, a new cutting-edge optical technique that received the Nobel Prize in Chemistry in 2014. In combination with innovative quantitative approaches and numerical simulations, they were also able to define the genome architecture at the nano-scale. Most importantly, they found that the nucleosomes are assembled in irregular groups across the chromatin and nucleosome-free-DNA regions separate these groups.
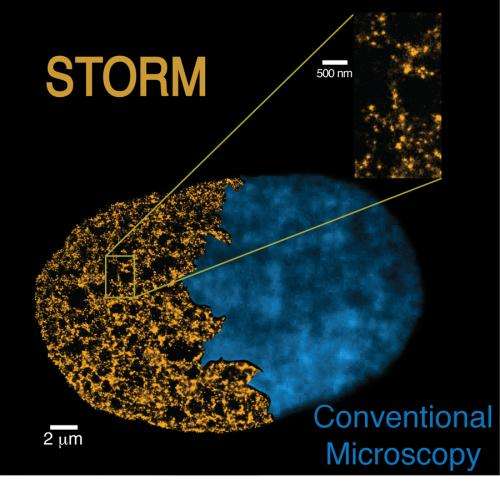
Biologists and physicists have been working together to take a step forward in chromatin fibre observations and studies. "By using the STORM technique, a new super-resolution microscopy method, we have been able to view and even count nucleosomes across the chromatin fibers and determine their organisation. STORM overcomes the diffraction limit that normally restricts the spatial resolution of conventional microscopes and enables us to precisely define the chromatin fibre structure", states Prof. Melike Lakadamyali, group leader at ICFO.
This enabling technique allowed the researchers to go deeper and, by comparing stem cells to differentiated cells (specialised cells that have already acquired their role), they observed key differences in the chromatin fibre architectures of both cells. Pia Cosma, group leader and ICREA research professor at the CRG explains, "We found that stem cells have a different chromatin structure than somatic (specialised) cells. At the same time, this difference correlates with the level of pluripotency. The more pluripotent a cell is, the less dense is its packaging. It gives us new clues to understand the stem cells functioning and their genomic structure, which will be helpful for example, in studying cell reprogramming".
What scientists have found is that DNA is not regularly packaged with nucleosomes, instead nucleosomes are assembled in groups of varying sizes, called "nucleosome clutches" -because of their similarity to egg clutches-. They found that pluripotent stem cells have, on average, clutches with less density of nucleosomes. In addition, clutch size is related to the pluripotency potential of stem cells, meaning that the more pluripotent a cell is, the less nucleosomes are included in its clutches.
Even though all the cells in our body have the same genetic information, they are not expressing all the genes at the same time. So, when a cell specialises, some of the DNA regions are silenced or less accessible to the molecule that reads the genome: the RNA polymerase. Depending on the specialisation of the cells, different levels of DNA packaging will occur. This new work published in the prestigious journal Cell, establishes a new understanding of how the chromatin fibre is assembled and packaged forming a specific DNA structure in every cell.
This research definitively contributes to the understanding of a novel feature of stem cells and their DNA structure, which is important for maintaining an induced pluripotent state. A joint patent has been filed by ICFO and CRG, who are now exploring business opportunities for marketing the classification of "stemness" state of cells, ie, their degree of pluripotency. This technique could determine with single cell sensitivity the pluripotency potential of stem cells, thus having the capacity of becoming a standard method of quality control of stem or pluripotent cells before their use in cell therapy or research in biomedicine.
More information: Ricci et al.: "Chromatin fibers are formed by heterogeneous groups of nucleosomes in vivo" Published in Cell on March 12 2015. Doi: dx.doi.org/10.1016/j.cell.2015.01.054
Journal information: Cell
Provided by ICFO-The Institute of Photonic Sciences