LSU researchers apply modern genomic analysis to historic bird collection
An international team of scientists has completed the largest whole genome study of a single class of animals to date. To map the tree of life for birds, the team sequenced, assembled and compared full genomes of 48 bird species representing all major branches of modern birds including ostrich, hummingbird, crow, duck, falcon, parrot, crane, ibis, woodpecker and eagle species. The researchers have been working on this ambitious genetic tree of life, or phylogeny, project for four years.
As part of the Avian Phylogenomics Consortium—comprised of more than 200 scientists from 80 institutions across 20 countries—LSU researchers' work was published today in three research papers featured in a special issue of Science.
"Mapping the tree of life is one of the grand challenges in biological science. Being part of this project is like being part of the writing of evolutionary history," said Robb Brumfield, director of the LSU Museum of Natural Science and Roy Paul Daniels Professor of Biological Sciences.
The LSU Museum of Natural Science maintains one of the world's largest frozen tissue collections of birds, reptiles, amphibians and mammals. Since 1979, the museum's curators and researchers have collected samples from unexplored regions of the world. They carry liquid nitrogen tanks in the field to flash freeze the tissue samples, which are brought back and stored in the Museum's Collection of Genetic Resources. They provided six of the 48 avian tissue samples whose genomes were sequenced for this project.
"When we collect a specimen, we try to preserve it in a way that will maximize its future utility. The challenge is that we don't know what the future uses will be," Brumfield said. "Ornithologists collecting bird tissues in 1979 had no idea that the entire genomes of these birds could one day be sequenced and featured in a study like this. It would have been impossible to envision."
One of the flagship papers published in Science titled, "Whole-genome analyses resolve early branches in the tree of life of modern birds," presents a well-resolved new tree of life for birds, based on whole-genome data. Several LSU scientists are co-authors on this paper including Brumfield, Assistant Professor of Biological Sciences Brant Faircloth, Curator of Genetic Resources Frederick Sheldon and former Museum of Natural Science Post-doctoral Researchers Elizabeth Derryberry and John McCormack.
This ambitious international research project is significant because it disentangles parts of the bird tree of life that have been particularly challenging to resolve. Because modern birds split into species early in time and in quick succession, the different lineages did not evolve enough distinct genetic differences at the genomic level to clearly determine their early branching order using smaller amounts of data, the researchers said. To overcome this problem and resolve the timing and relationships of modern birds, the consortium authors used whole-genome sequences to infer the bird species tree.
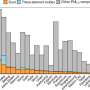
LSU's Brant Faircloth contributed to the reconstruction of the bird tree by running statistical analyses of genomic loci called ultraconserved elements, or UCEs that are found throughout the genomes of many organisms. His group, in collaboration with Brumfield's students and post-doctoral researchers, developed the technique for studying UCEs. By aligning these highly conserved parts of the genome and analyzing the subtle variations across the different species, the UCE data helped piece together this new tree of life for birds. Faircloth found that the UCE data provided consistent results to infer evolutionary history whereas traditionally, scientists have used data from the protein coding parts of the genome called exons for this purpose.
"It opens up the potential in terms of the types of data people will collect and use in the future when reconstructing the tree of life," Faircloth said.
Once they inferred a tree from the data, collaborators used the fossil record to date the divergence of each species across millions of years. The result is a genome-scale phylogeny of birds that dates the evolutionary expansion of modern birds, or Neoaves, to 66 million years ago, when a mass extinction event killed off the dinosaurs, but a few bird lineages survived.
In addition to resolving the early branches of Neoaves, this new work supports conclusions about some relationships that have been long-debated. For example, the findings support three independent origins of waterbirds. They also indicate that the common ancestor of land birds, including songbirds, parrots, woodpeckers, owls, eagles and falcons, was an apex predator, which also gave rise to the giant terror birds that once roamed the Americas.
The second flagship paper titled, "Comparative genomics reveals insights into avian genome evolution and adaptation," illustrates that genomic diversity across birds correlates with adaptation to different lifestyles and evolutionary traits. These findings help bridge the chasm between micro and macroevolution.
Lastly, a third paper titled, "Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs," was co-authored by Faircloth, McCormack and fellow former LSU Post-doctoral Researcher David Ray. They used newly assembled genomes of the closest living relatives to birds—crocodilians —to provide context for the diversification of archosaurs, a group that includes birds, crocodilians and dinosaurs. Using the genomes of the American alligator, the saltwater crocodile and the Indian gharial, the scientists showed that crocodilian genomes are among the slowest evolving vertebrates. Their collaborators also used these genomes to infer the genome sequence of the common ancestor of archosaurs and therefore all dinosaurs including those that became extinct 66 million years ago. Tissues of the individual crocodilians whose genomes were sequenced by the team will also be stored in the LSU Museum of Natural Science Collection of Genetic Resources.
Setting up the pipeline for the large-scale study of whole genomes—collecting and organizing tissue samples; extracting the DNA; analyzing its quality; sequencing, managing and analyzing torrents of new data—has been a massive undertaking. But the scientists say their work should help inform other major efforts for the comprehensive sequencing of vertebrate classes. To encourage other researchers to dig through this "big data" and discover new patterns that were not seen in small-scale data before, the Avian Genome Consortium has released the full dataset to the public in GigaScience and other databases.
"The next grand challenge in our field would be to expand upon this and sequence the genomes of all birds," Brumfield said.
More information: phys.org/news/2014-12-internat … -bird-evolution.html
Journal information: Science , GigaScience
Provided by Louisiana State University