Scientists call for investigation of mysterious cloud-like collections in cells
About 50 years ago, electron microscopy revealed the presence of tiny blob-like structures that form inside cells, move around and disappear. But scientists still don't know what they do—even though these shifting cloud-like collections of proteins are believed to be crucial to the life of a cell, and therefore could offer a new approach to disease treatment.
In the Journal of Cell Biology, two researchers are issuing a call to investigators from various backgrounds, from biophysics to cell biology, to focus their attention on the role of these formations— for which they coin a new unifying term "assemblages."
"I want to know what these assemblages are doing in Ewing sarcoma, the disease I concentrate on—and I would think all other researchers who study human biology would want to know their functions in both health and disease," says Jeffrey Toretsky, MD, professor in the department of oncology and pediatrics at Georgetown Lombardi Comprehensive Cancer Center.
So Toretsky partnered with co-author Peter Wright, PhD, professor in the department of integrative structural and computational biology at The Scripps Research Institute in La Jolla, Calif., to pull together all the biophysics and protein biochemistry knowledge available on assemblages into a review article. Toretsky also called on the expertise of chemists and physicists from Georgetown University.
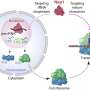
The authors say these assemblages are often, but not always, made up of proteins that are intrinsically disordered, meaning that they do not assume a specific shape in order to fit like a lock and key onto other proteins. These intrinsically disordered proteins seem to find each other and then form into gel-like assemblages—a process called "phase separation"—that can trap and interact with other proteins and even RNA, biological molecules that help decode and regulate genes.
When their work is done—whatever that is—the assemblages dissolve, Toretsky says.
"It is only in the last five years that researchers have begun recognizing that proteins without fixed structures may have important transitional properties that change based upon their local abundance in cells," he says.
Toretsky suspects that if these assemblages play a role in disease, they could be targeted with a small molecule. "Current drug-discovery dogma suggests that it is very hard to make a small molecule to prevent two structured proteins from interacting. However, small molecules have a greater likelihood of disrupting intrinsically disordered protein-protein interactions," he says.
"This review links together very basic biologic phenomena of protein interaction with the potential for new drug discovery," Toretsky says. "It's an exciting challenge."
Journal information: Journal of Cell Biology
Provided by Georgetown University Medical Center