Interactions of Earth's smallest players have global impact

A new study reveals the interactions among bacteria and viruses that prey on them thriving in oxygen minimum zones—stretches of ocean starved for oxygen that occur around the globe. Understanding such microbial communities in their natural environments is an important step in understanding global processes, including climate.
A complex web of interaction among viruses, bacteria and their environment is becoming ever more untangled by a growing international collaboration between Matthew Sullivan, associate professor in the University of Arizona's Department of Ecology and Evolutionary Biology, and Steven Hallam from the University of British Columbia in Canada.
"Bacteria are drivers of nutrient and energy cycles that power the earth," said Sullivan, who is also a member of the UA's BIO5 Institute. "As the climate is changing, so are the environments these bacteria live in, and they in turn loop back to impact their environments. Viruses are also in the mix mediating microbial processes, but how and to what extent? Those are the questions we are trying to answer."
The latest chapter in understanding this complex ecosystem was led by Simon Roux, a post-doctoral research scientist in Sullivan's lab. In a study published in a new, nonprofit, open-access journal called eLife, Roux focused on viruses and the microbes they infect in the northeast Pacific Ocean, specifically in vast stretches of open ocean depleted of oxygen, which scientists call oxygen minimum zones, or OMZs.
"There are, of course, areas with less oxygen scattered throughout the world's oceans," Sullivan said. "However, the more worrisome OMZs form when oxygen concentrations become limiting to life due to physical processes preventing gas exchange with the atmosphere and microbes respiring to draw down available oxygen."
Sullivan explained that as water columns become increasingly stratified – in other words, no longer mix well – OMZs are expanding and intensifying due to microbial activity that drives chemical transformations resulting in loss of fixed nitrogen that phytoplankton depends on; accumulation of hydrogen sulfide, a chemical toxic to animals and plants; and production of greenhouse gases.

The eLife study focused on a sulfur-oxidizing bacterium called SUP05, which is dominant in OMZs and, like most microbes in the "wild," has not been successfully cultivated in a lab setting. Using cutting-edge genome sequencing technology, the genomes of 127 single cells were generated and then screened using novel computational approaches to look for virus DNA in these single-cell genomes.
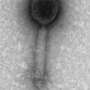
Also called bacteriophages, viruses that prey on bacteria are less familiar to most people than their flu- or cold-causing cousins, but they control processes of global importance. For example, they determine how much oxygen goes from the oceans into the atmosphere in exchange for carbon dioxide, they influence climate patterns across the Earth and they alter the assemblages of microorganisms competing in the environment.
"The last couple of years have seen a spectacular development of the single-cell genome technique, in which scientists are able to isolate one single cell and sequence all of its genetic material," Roux said. "Originally used to sequence the genome of organisms without having to cultivate them in the lab, the resulting datasets are also extremely interesting from a virus point-of-view to be able to clearly identify and directly link virus to host."
From these 127 single-cell genomes, 69 new SUP05 viruses were identified representing five viral groups not yet known to science. These new genomes were then used as references to query 186 available environmental microbial and viral sequence datasets to show that these new viruses were persistent over time and largely did not occur outside the study site. In other words, these viruses and their hosts thrive in oxygen-depleted parts of the ocean where not much else can survive, and they stay there.
Not only did this study reveal the type of viruses that infect these microbes but also features of how these viruses and hosts interact. For one, many SUP05 cells are commonly infected by viruses in nature—about one in three. Second, these SUP05 viruses encode metabolic genes that impact the very functions that these SUP05 bacteria contribute to the OMZ ecosystem—energy and nutrient cycling.
"The finding that viruses could directly manipulate SUP05 sulfur cycling that drives its coupled carbon fixation is critical to developing predictive models about microbial processes in the dark ocean," Sullivan said. "However, it is becoming less and less surprising as we have known for a decade that viruses encode core photosynthesis genes when they infect photosynthetic cyanobacteria."
The work published in the eLife journal is unprecedented in its breadth and scope and represents some of the first population-based viral ecology in natural systems. Notably, the effort to develop the tools needed to establish a population-based viral ecology is a central theme in Sullivan's research, resulting in the recent invention of a viral tagging method that was published in Nature.
Roux and Sullivan are affiliated with the newly launched UA Ecosystem Genomics Institute, led by Scott Saleska, associate professor in ecology and evolutionary biology, which seeks to use modern approaches to build an ecosystems-level understanding of microbial communities in nature. Seed funding for EGI was provided by the UA Technology and Research Initiative Fund through the Water, Environmental and Energy Solutions initiative.
"This study represents a solid first step at doing so in an ocean ecosystem focused on OMZs, whose recent expansion we hope to see reversed," Sullivan said.
Provided by University of Arizona