Molecular gate that could keep cancer cells locked up
In a study published today in Genes & Development, Dr Christian Speck from the MRC Clinical Sciences Centre's DNA Replication group, in collaboration with Brookhaven National Laboratory (BNL), New York, reveal the intricate mechanisms involved in the enzyme that governs DNA duplication during cell division. By developing a sophisticated system using synthetic, chemical and structural biology approaches, the study reveals how a key enzyme involved in duplicating genetic information embraces DNA through a gated system, which opens up at precise positions allowing for a highly regulated replication process. This work enhances current understanding of an essential biological process and suggests a route for stopping cell division in disease such as cancer.
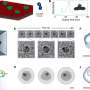
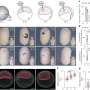
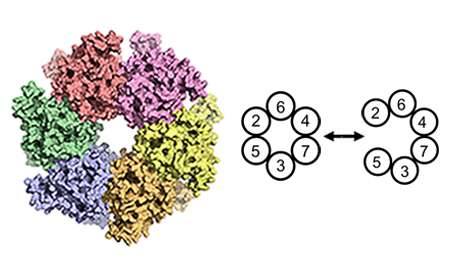
When a cell divides, genetic information is duplicated in a process known as DNA replication. For this to occur, a 'replication machine' is assembled on top of the DNA prior to duplication. A protein complex known as ORC that recognises the DNA replication origin initiates the whole process. Next, an enzyme, MCM2-7 helicase, whose role is to unwind and separate the two strands of the DNA helix, is loaded onto the DNA by the machine system ORC. The helicase is a ring shaped enzyme composed of six subunits (hexamer), though how the ring structure opens and encircles the DNA has, until now, remained a mystery.
Initial theories within in field assumed the helicase to exist in an open ring conformation. Speck's team argued that this would undoubtedly lead to poorly regulated DNA replication with no control or specificity. To examine the helicase activity in more detail, Jingchuan Sun at BNL used an electron microscope and revealed, contrary to initial theories, the helicase actually existed as a closed ring conformation.
To pinpoint where within the six subunits, the helicase opens to encompass the DNA, the team generated linkages that blocked ring opening at various positions. They found that if they blocked one specific interface, between MCM2 and MCM5, DNA could not enter. A small molecule called rapamycin brings the linkages together; such a molecular switch can be used to control DNA entry to the MCM ring and subsequent DNA replication. "Both in the context of our in vitro and in vivo experiments, we showed that opening of the MCM2/MCM5 interface is essential for helicase loading onto DNA," explains Christian.
"The field has known for a while that DNA can pass into the MCM2-7 ring, but has never been sure which MCM subunits are used for regulated helicase loading. By designing an elegant experiment, the Speck laboratory has now shown once and for all that the MCM2-5 is the only DNA entry point," says collaborator Huilin Li at BNL.
In eukaryotes, the MCM2-7 helicase forms a double hexamer (with another MCM2-7 unit) when it is loaded onto DNA. In this study, the group also settled the longstanding dispute surrounding whether the helicase is actually loaded as a single hexamer, which then dimerises, or is loaded as a dimer at the offset. They concluded that the helicase is in fact loaded as a single hexamer before forming a double hexamer.
In a successful collaboration that harnesses the electron microscopy expertise at BNL with the chemical biology and genetic expertise at the MRC Clinical Sciences Centre, the study addresses key questions detailing the processes involved in DNA replication. "Our work is aimed at understanding the molecular mechanism of DNA replication at a fundamental level. Yet our findings could also have important implications, possibly pointing to new ways to fight cancer, because DNA duplication is a prime target to inhibit cancer cell growth," says Christian.
More information: Genes & Development, 28 (15)
Journal information: Genes & Development
Provided by MRC Clinical Sciences Centre